Convergent evolution in structural elements of proteins investigated using cross profile analysis
- PMID: 22244085
- PMCID: PMC3398312
- DOI: 10.1186/1471-2105-13-11
Convergent evolution in structural elements of proteins investigated using cross profile analysis
Abstract
Background: Evolutionary relations of similar segments shared by different protein folds remain controversial, even though many examples of such segments have been found. To date, several methods such as those based on the results of structure comparisons, sequence-based classifications, and sequence-based profile-profile comparisons have been applied to identify such protein segments that possess local similarities in both sequence and structure across protein folds. However, to capture more precise sequence-structure relations, no method reported to date combines structure-based profiles, and sequence-based profiles based on evolutionary information. The former are generally regarded as representing the amino acid preferences at each position of a specific conformation of protein segment. They might reflect the nature of ancient short peptide ancestors, using the results of structural classifications of protein segments.
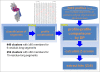
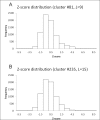
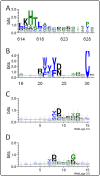
Results: This report describes the development and use of "Cross Profile Analysis" to compare sequence-based profiles and structure-based profiles based on amino acid occurrences at each position within a protein segment cluster. Using systematic cross profile analysis, we found structural clusters of 9-residue and 15-residue segments showing remarkably strong correlation with particular sequence profiles. These correlations reflect structural similarities among constituent segments of both sequence-based and structure-based profiles. We also report previously undetectable sequence-structure patterns that transcend protein family and fold boundaries, and present results of the conformational analysis of the deduced peptide of a segment cluster. These results suggest the existence of ancient short-peptide ancestors.
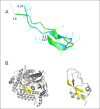
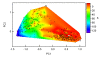
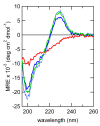
Conclusions: Cross profile analysis reveals the polyphyletic and convergent evolution of β-hairpin-like structures, which were verified both experimentally and computationally. The results presented here give us new insights into the evolution of short protein segments.
Figures
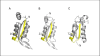


References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

