A comprehensive dataset of genes with a loss-of-function mutant phenotype in Arabidopsis
- PMID: 22247268
- PMCID: PMC3291275
- DOI: 10.1104/pp.111.192393
A comprehensive dataset of genes with a loss-of-function mutant phenotype in Arabidopsis
Abstract
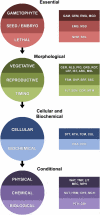
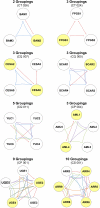
Despite the widespread use of Arabidopsis (Arabidopsis thaliana) as a model plant, a curated dataset of Arabidopsis genes with mutant phenotypes remains to be established. A preliminary list published nine years ago in Plant Physiology is outdated, and genome-wide phenotype information remains difficult to obtain. We describe here a comprehensive dataset of 2,400 genes with a loss-of-function mutant phenotype in Arabidopsis. Phenotype descriptions were gathered primarily from manual curation of the scientific literature. Genes were placed into prioritized groups (essential, morphological, cellular-biochemical, and conditional) based on the documented phenotypes of putative knockout alleles. Phenotype classes (e.g. vegetative, reproductive, and timing, for the morphological group) and subsets (e.g. flowering time, senescence, circadian rhythms, and miscellaneous, for the timing class) were also established. Gene identities were classified as confirmed (through molecular complementation or multiple alleles) or not confirmed. Relationships between mutant phenotype and protein function, genetic redundancy, protein connectivity, and subcellular protein localization were explored. A complementary dataset of 401 genes that exhibit a mutant phenotype only when disrupted in combination with a putative paralog was also compiled. The importance of these genes in confirming functional redundancy and enhancing the value of single gene datasets is discussed. With further input and curation from the Arabidopsis community, these datasets should help to address a variety of important biological questions, provide a foundation for exploring the relationship between genotype and phenotype in angiosperms, enhance the utility of Arabidopsis as a reference plant, and facilitate comparative studies with model genetic organisms.
Figures





References
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al. (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657 - PubMed
-
- Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815 - PubMed
-
- Becraft PW, Li K, Dey N, Asuncion-Crabb Y. (2002) The maize dek1 gene functions in embryonic pattern formation and cell fate specification. Development 129: 5217–5225 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

