Host gene targets for novel influenza therapies elucidated by high-throughput RNA interference screens
- PMID: 22247330
- PMCID: PMC3316894
- DOI: 10.1096/fj.11-193466
Host gene targets for novel influenza therapies elucidated by high-throughput RNA interference screens
Abstract
Influenza virus encodes only 11 viral proteins but replicates in a broad range of avian and mammalian species by exploiting host cell functions. Genome-wide RNA interference (RNAi) has proven to be a powerful tool for identifying the host molecules that participate in each step of virus replication. Meta-analysis of findings from genome-wide RNAi screens has shown influenza virus to be dependent on functional nodes in host cell pathways, requiring a wide variety of molecules and cellular proteins for replication. Because rapid evolution of the influenza A viruses persistently complicates the effectiveness of vaccines and therapeutics, a further understanding of the complex host cell pathways coopted by influenza virus for replication may provide new targets and strategies for antiviral therapy. RNAi genome screening technologies together with bioinformatics can provide the ability to rapidly identify specific host factors involved in resistance and susceptibility to influenza virus, allowing for novel disease intervention strategies.
Figures
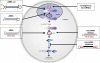
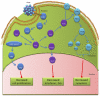
References
-
- Daszak P., Cunningham A. A., Hyatt A. D. (2000) Emerging infectious diseases of wildlife–threats to biodiversity and human health. Science 287, 443–449 - PubMed
-
- World Health Organization (2009) Overview of pandemic (H1N1) 2009 in the WHO European Region. Retrieved March 11, 2010, from http://www.euro.who.int/influenza/AH1N1/20091026_1
-
- World Health Organization (2009) Seasonal influenza. Retrieved March 10, 2010, from http://www.euro.who.int/influenza/20080618_1
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

