The organisation of Ebola virus reveals a capacity for extensive, modular polyploidy
- PMID: 22247782
- PMCID: PMC3256159
- DOI: 10.1371/journal.pone.0029608
The organisation of Ebola virus reveals a capacity for extensive, modular polyploidy
Abstract
Background: Filoviruses, including Ebola virus, are unusual in being filamentous animal viruses. Structural data on the arrangement, stoichiometry and organisation of the component molecules of filoviruses has until now been lacking, partially due to the need to work under level 4 biological containment. The present study provides unique insights into the structure of this deadly pathogen.
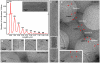
Methodology and principal findings: We have investigated the structure of Ebola virus using a combination of cryo-electron microscopy, cryo-electron tomography, sub-tomogram averaging, and single particle image processing. Here we report the three-dimensional structure and architecture of Ebola virus and establish that multiple copies of the RNA genome can be packaged to produce polyploid virus particles, through an extreme degree of length polymorphism. We show that the helical Ebola virus inner nucleocapsid containing RNA and nucleoprotein is stabilized by an outer layer of VP24-VP35 bridges. Elucidation of the structure of the membrane-associated glycoprotein in its native state indicates that the putative receptor-binding site is occluded within the molecule, while a major neutralizing epitope is exposed on its surface proximal to the viral envelope. The matrix protein VP40 forms a regular lattice within the envelope, although its contacts with the nucleocapsid are irregular.
Conclusions: The results of this study demonstrate a modular organization in Ebola virus that accommodates a well-ordered, symmetrical nucleocapsid within a flexible, tubular membrane envelope.
Conflict of interest statement
Figures





References
-
- Caspar DL, Klug A. Physical principles in the construction of regular viruses. Cold Spring Harb Symp Quant Biol. 1962;27:1–24. - PubMed
-
- Hosaka Y, Kitano H, Ikeguchi S. Studies on the pleomorphism of HVJ virons. Virology. 1966;29:205–221. - PubMed
-
- Noda T, Sagara H, Yen A, Takada A, Kida H, et al. Architecture of ribonucleoprotein complexes in influenza A virus particles. Nature. 2006;439:490–492. - PubMed
-
- Geisbert TW, Hensley LE. Ebola virus: new insights into disease aetiopathology and possible therapeutic interventions. Expert Rev Mol Med. 2004;6:1–24. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

