A high throughput scintillation proximity imaging assay for protein methyltransferases
- PMID: 22256970
- PMCID: PMC3553658
- DOI: 10.2174/138620712800194468
A high throughput scintillation proximity imaging assay for protein methyltransferases
Abstract
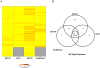
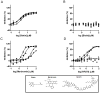
Protein methyltransferases (PMTs) orchestrate epigenetic modifications through post-translational methylation of various protein substrates including histones. Since dysregulation of this process is widely implicated in many cancers, it is of pertinent interest to screen inhibitors of PMTs, as they offer novel target-based opportunities to discover small molecules with potential chemotherapeutic use. We have thus developed an enzymatic screening strategy, which can be adapted to scintillation proximity imaging assay (SPIA) format, to identify these inhibitors. We took advantage of S-adenosyl-L-[3H-methyl]-methionine availability and monitored the enzymatically catalyzed [3H]-methyl addition on lysine residues of biotinylated peptide substrates. The radiolabeled peptides were subsequently captured by streptavidin coated SPA imaging PS beads. We applied this strategy to four PMTs: SET7/9, SET8, SETD2, and EuHMTase1, and optimized assay conditions to achieve Z' values ranging from 0.48 to 0.91. The robust performance of this SPIA for the four PMTs was validated in a pilot screen of approximately 7,000 compounds. We identified 80 cumulative hits across the four targets. NF279, a suramin analogue, was found to specifically inhibit SET7/9 and SETD2 with IC50 values of 1.9 and 1.1 μM, respectively. Another identified compound, Merbromin, a topical antiseptic, was classified as a pan-active inhibitor of the four PMTs. These findings demonstrate that our proposed SPIA strategy is generic for multiple PMTs and can be successfully implemented to identify novel and specific inhibitors of PMTs. The specific PMT inhibitors may constitute a new class of anti-proliferative agents for potential therapeutic use.
Figures
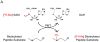
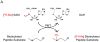
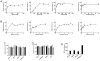



Similar articles
-
Small-molecule inhibitors of the protein methyltransferase SET7/9 identified in a high-throughput screen.J Biomol Screen. 2012 Sep;17(8):1102-9. doi: 10.1177/1087057112452137. Epub 2012 Jul 5. J Biomol Screen. 2012. PMID: 22772057
-
Formulating a fluorogenic assay to evaluate S-adenosyl-L-methionine analogues as protein methyltransferase cofactors.Mol Biosyst. 2011 Nov;7(11):2970-81. doi: 10.1039/c1mb05230f. Epub 2011 Aug 24. Mol Biosyst. 2011. PMID: 21866297 Free PMC article.
-
A scintillation proximity assay for histone demethylases.Anal Biochem. 2014 Oct 15;463:54-60. doi: 10.1016/j.ab.2014.06.023. Epub 2014 Jul 8. Anal Biochem. 2014. PMID: 25010373
-
Inhibitors of protein methyltransferases as chemical tools.Epigenomics. 2015;7(8):1327-38. doi: 10.2217/epi.15.87. Epub 2015 Dec 8. Epigenomics. 2015. PMID: 26646500 Free PMC article. Review.
-
Recent progress in developing selective inhibitors of protein methyltransferases.Curr Opin Chem Biol. 2017 Aug;39:100-108. doi: 10.1016/j.cbpa.2017.06.013. Epub 2017 Jun 26. Curr Opin Chem Biol. 2017. PMID: 28662389 Free PMC article. Review.
Cited by
-
Kinetic isotope effects reveal early transition state of protein lysine methyltransferase SET8.Proc Natl Acad Sci U S A. 2016 Dec 27;113(52):E8369-E8378. doi: 10.1073/pnas.1609032114. Epub 2016 Dec 9. Proc Natl Acad Sci U S A. 2016. PMID: 27940912 Free PMC article.
-
Targeted drug combination therapy design based on driver genes.Oncotarget. 2019 Sep 3;10(51):5255-5266. doi: 10.18632/oncotarget.26985. eCollection 2019 Sep 3. Oncotarget. 2019. PMID: 31523388 Free PMC article.
-
Chemical & RNAi screening at MSKCC: a collaborative platform to discover & repurpose drugs to fight disease.Comb Chem High Throughput Screen. 2014 May;17(4):298-318. doi: 10.2174/1386207317666140323132222. Comb Chem High Throughput Screen. 2014. PMID: 24661215 Free PMC article.
-
Methionine Adenosyltransferase Engineering to Enable Bioorthogonal Platforms for AdoMet-Utilizing Enzymes.ACS Chem Biol. 2020 Mar 20;15(3):695-705. doi: 10.1021/acschembio.9b00943. Epub 2020 Mar 3. ACS Chem Biol. 2020. PMID: 32091873 Free PMC article.
-
The dynamic conformational landscape of the protein methyltransferase SETD8.Elife. 2019 May 13;8:e45403. doi: 10.7554/eLife.45403. Elife. 2019. PMID: 31081496 Free PMC article.
References
-
- Kouzarides T. Chromatin modifications and their function. Cell. 2007;128(4):693–705. - PubMed
-
- Copeland RA, Solomon ME, Richon VM. Protein methyltransferases as a target class for drug discovery. Nat Rev Drug Discov. 2009;8:724–732. - PubMed
-
- Suh-Lailam BB, Hevel JM. A fast and efficient method for quantitative measurement of S-adenosyl-L-methionine-dependent methyltransferase activity with protein substrates. Anal Biochem. 2010;398:218–224. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

