Selective subversion of autophagy complexes facilitates completion of the Brucella intracellular cycle
- PMID: 22264511
- PMCID: PMC3266535
- DOI: 10.1016/j.chom.2011.12.002
Selective subversion of autophagy complexes facilitates completion of the Brucella intracellular cycle
Abstract
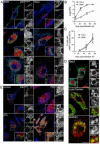
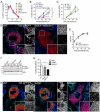
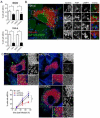
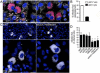
Autophagy is a cellular degradation process that can capture and eliminate intracellular microbes by delivering them to lysosomes for destruction. However, pathogens have evolved mechanisms to subvert this process. The intracellular bacterium Brucella abortus ensures its survival by forming the Brucella-containing vacuole (BCV), which traffics from the endocytic compartment to the endoplasmic reticulum (ER), where the bacterium proliferates. We show that Brucella replication in the ER is followed by BCV conversion into a compartment with autophagic features (aBCV). While Brucella trafficking to the ER was unaffected in autophagy-deficient cells, aBCV formation required the autophagy-initiation proteins ULK1, Beclin 1, and ATG14L and PI3-kinase activity. However, aBCV formation was independent of the autophagy-elongation proteins ATG5, ATG16L1, ATG4B, ATG7, and LC3B. Furthermore, aBCVs were required to complete the intracellular Brucella lifecycle and for cell-to-cell spreading, demonstrating that Brucella selectively co-opts autophagy-initiation complexes to subvert host clearance and promote infection.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures


Comment in
-
Brucella "hitches a ride" with autophagy.Cell Host Microbe. 2012 Jan 19;11(1):2-4. doi: 10.1016/j.chom.2012.01.003. Cell Host Microbe. 2012. PMID: 22264507
-
Cellular microbiology: an unconventional exit for Brucella.Nat Rev Microbiol. 2012 Feb 6;10(3):160. doi: 10.1038/nrmicro2749. Nat Rev Microbiol. 2012. PMID: 22307031 No abstract available.
Similar articles
-
Brucella "hitches a ride" with autophagy.Cell Host Microbe. 2012 Jan 19;11(1):2-4. doi: 10.1016/j.chom.2012.01.003. Cell Host Microbe. 2012. PMID: 22264507
-
Replication of Brucella abortus and Brucella melitensis in fibroblasts does not require Atg5-dependent macroautophagy.BMC Microbiol. 2014 Sep 2;14:223. doi: 10.1186/s12866-014-0223-5. BMC Microbiol. 2014. PMID: 25179110 Free PMC article.
-
A T4SS Effector Targets Host Cell Alpha-Enolase Contributing to Brucella abortus Intracellular Lifestyle.Front Cell Infect Microbiol. 2016 Nov 16;6:153. doi: 10.3389/fcimb.2016.00153. eCollection 2016. Front Cell Infect Microbiol. 2016. PMID: 27900285 Free PMC article.
-
Invasion and intracellular trafficking of Brucella abortus in nonphagocytic cells.Microbes Infect. 2000 Jun;2(7):829-35. doi: 10.1016/s1286-4579(00)90368-x. Microbes Infect. 2000. PMID: 10955964 Review.
-
Autophagy Evasion and Endoplasmic Reticulum Subversion: The Yin and Yang of Legionella Intracellular Infection.Annu Rev Microbiol. 2016 Sep 8;70:413-33. doi: 10.1146/annurev-micro-102215-095557. Annu Rev Microbiol. 2016. PMID: 27607556 Review.
Cited by
-
The Mechanism of Facultative Intracellular Parasitism of Brucella.Int J Mol Sci. 2021 Apr 1;22(7):3673. doi: 10.3390/ijms22073673. Int J Mol Sci. 2021. PMID: 33916050 Free PMC article. Review.
-
Crosstalk between autophagy and inflammatory signalling pathways: balancing defence and homeostasis.Nat Rev Immunol. 2016 Nov;16(11):661-675. doi: 10.1038/nri.2016.100. Epub 2016 Oct 3. Nat Rev Immunol. 2016. PMID: 27694913 Free PMC article. Review.
-
Evasion of host defense by Brucella.Cell Insight. 2023 Dec 17;3(1):100143. doi: 10.1016/j.cellin.2023.100143. eCollection 2024 Feb. Cell Insight. 2023. PMID: 38250017 Free PMC article. Review.
-
Immunosuppressive Mechanisms in Brucellosis in Light of Chronic Bacterial Diseases.Microorganisms. 2022 Jun 21;10(7):1260. doi: 10.3390/microorganisms10071260. Microorganisms. 2022. PMID: 35888979 Free PMC article. Review.
-
Entrance and survival of Brucella pinnipedialis hooded seal strain in human macrophages and epithelial cells.PLoS One. 2013 Dec 20;8(12):e84861. doi: 10.1371/journal.pone.0084861. eCollection 2013. PLoS One. 2013. PMID: 24376851 Free PMC article.
References
-
- Archambaud C, Salcedo SP, Lelouard H, Devilard E, de Bovis B, Van Rooijen N, Gorvel JP, Malissen B. Contrasting roles of macrophages and dendritic cells in controlling initial pulmonary Brucella infection. Eur J Immunol. 2010;40:3458–3471. - PubMed
-
- Arellano-Reynoso B, Lapaque N, Salcedo S, Briones G, Ciocchini AE, Ugalde R, Moreno E, Moriyon I, Gorvel JP. Cyclic beta-1,2-glucan is a brucella virulence factor required for intracellular survival. Nat Immunol. 2005;6:618–625. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

