The transcriptional activator ZNF143 is essential for normal development in zebrafish
- PMID: 22268977
- PMCID: PMC3282657
- DOI: 10.1186/1471-2199-13-3
The transcriptional activator ZNF143 is essential for normal development in zebrafish
Abstract
Background: ZNF143 is a sequence-specific DNA-binding protein that stimulates transcription of both small RNA genes by RNA polymerase II or III, or protein-coding genes by RNA polymerase II, using separable activating domains. We describe phenotypic effects following knockdown of this protein in developing Danio rerio (zebrafish) embryos by injection of morpholino antisense oligonucleotides that target znf143 mRNA.
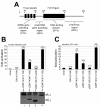
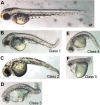
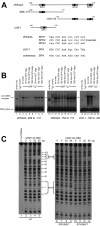
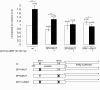
Results: The loss of function phenotype is pleiotropic and includes a broad array of abnormalities including defects in heart, blood, ear and midbrain hindbrain boundary. Defects are rescued by coinjection of synthetic mRNA encoding full-length ZNF143 protein, but not by protein lacking the amino-terminal activation domains. Accordingly, expression of several marker genes is affected following knockdown, including GATA-binding protein 1 (gata1), cardiac myosin light chain 2 (cmlc2) and paired box gene 2a (pax2a). The zebrafish pax2a gene proximal promoter contains two binding sites for ZNF143, and reporter gene transcription driven by this promoter in transfected cells is activated by this protein.
Conclusions: Normal development of zebrafish embryos requires ZNF143. Furthermore, the pax2a gene is probably one example of many protein-coding gene targets of ZNF143 during zebrafish development.
Figures


Similar articles
-
The Oct4-related PouV gene, pou5f3, mediates isthmus development in zebrafish by directly and dynamically regulating pax2a.Cells Dev. 2024 Sep;179:203933. doi: 10.1016/j.cdev.2024.203933. Epub 2024 Jun 20. Cells Dev. 2024. PMID: 38908828
-
Depletion of zebrafish essential and regulatory myosin light chains reduces cardiac function through distinct mechanisms.Cardiovasc Res. 2008 Jul 1;79(1):97-108. doi: 10.1093/cvr/cvn073. Epub 2008 Mar 14. Cardiovasc Res. 2008. PMID: 18343897 Free PMC article.
-
MiR-144 regulates hematopoiesis and vascular development by targeting meis1 during zebrafish development.Int J Biochem Cell Biol. 2014 Apr;49:53-63. doi: 10.1016/j.biocel.2014.01.005. Epub 2014 Jan 18. Int J Biochem Cell Biol. 2014. PMID: 24448023
-
Zebrafish pax8 is required for otic placode induction and plays a redundant role with Pax2 genes in the maintenance of the otic placode.Development. 2005 Jan;132(2):371-82. doi: 10.1242/dev.01587. Epub 2004 Dec 16. Development. 2005. PMID: 15604103
-
Manipulating Galectin Expression in Zebrafish (Danio rerio).Methods Mol Biol. 2022;2442:425-443. doi: 10.1007/978-1-0716-2055-7_23. Methods Mol Biol. 2022. PMID: 35320539 Review.
Cited by
-
Two paralogous znf143 genes in zebrafish encode transcriptional activator proteins with similar functions but expressed at different levels during early development.BMC Mol Cell Biol. 2020 Jan 22;21(1):3. doi: 10.1186/s12860-020-0247-7. BMC Mol Cell Biol. 2020. PMID: 31969120 Free PMC article.
-
ZNF143 protein is an important regulator of the myeloid transcription factor C/EBPα.J Biol Chem. 2017 Nov 17;292(46):18924-18936. doi: 10.1074/jbc.M117.811109. Epub 2017 Sep 12. J Biol Chem. 2017. PMID: 28900037 Free PMC article.
-
ZNF143 is a transcriptional regulator of nuclear-encoded mitochondrial genes that acts independently of looping and CTCF.Mol Cell. 2025 Jan 2;85(1):24-41.e11. doi: 10.1016/j.molcel.2024.11.031. Epub 2024 Dec 20. Mol Cell. 2025. PMID: 39708805 Free PMC article.
-
A Mutation in ZNF143 as a Novel Candidate Gene for Endothelial Corneal Dystrophy.J Clin Med. 2019 Aug 6;8(8):1174. doi: 10.3390/jcm8081174. J Clin Med. 2019. PMID: 31390831 Free PMC article.
-
ZNF143 provides sequence specificity to secure chromatin interactions at gene promoters.Nat Commun. 2015 Feb 3;2:6186. doi: 10.1038/ncomms7186. Nat Commun. 2015. PMID: 25645053 Free PMC article.
References
-
- Myslinski E, Krol A, Carbon P. ZNF76 and ZNF143 Are Two Human Homologs of the Transcriptional Activator Staf. J Biol Chem. 1998;273:21998–22006. - PubMed
-
- Myslinski E, Gerard MA, Krol A, Carbon P. A Genome Scale Location Analysis of Human Staf/ZNF143-binding Sites Suggests a Widespread Role for Human Staf/ZNF143 in Mammalian Promoters. J Biol Chem. 2006;281:39953–39962. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

