The Ras protein superfamily: evolutionary tree and role of conserved amino acids
- PMID: 22270915
- PMCID: PMC3265948
- DOI: 10.1083/jcb.201103008
The Ras protein superfamily: evolutionary tree and role of conserved amino acids
Erratum in
- J Cell Biol. 2012 Feb 20;196(4):545
Abstract
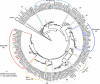
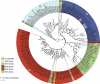
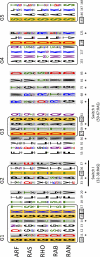
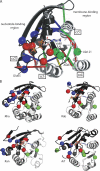
The Ras superfamily is a fascinating example of functional diversification in the context of a preserved structural framework and a prototypic GTP binding site. Thanks to the availability of complete genome sequences of species representing important evolutionary branch points, we have analyzed the composition and organization of this superfamily at a greater level than was previously possible. Phylogenetic analysis of gene families at the organism and sequence level revealed complex relationships between the evolution of this protein superfamily sequence and the acquisition of distinct cellular functions. Together with advances in computational methods and structural studies, the sequence information has helped to identify features important for the recognition of molecular partners and the functional specialization of different members of the Ras superfamily.
Figures

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

