Application of RNA-Seq transcriptome analysis: CD151 is an Invasion/Migration target in all stages of epithelial ovarian cancer
- PMID: 22272937
- PMCID: PMC3288733
- DOI: 10.1186/1757-2215-5-4
Application of RNA-Seq transcriptome analysis: CD151 is an Invasion/Migration target in all stages of epithelial ovarian cancer
Abstract
Background: RNA-Seq allows a theoretically unbiased analysis of both genome-wide transcription levels and mutation status of a tumor. Using this technique we sought to identify novel candidate therapeutic targets expressed in epithelial ovarian cancer (EOC).
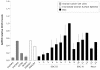
Methods: Specifically, we sought candidate invasion/migration targets based on expression levels across all tumors, novelty of expression in EOC, and known function. RNA-Seq analysis revealed the high expression of CD151, a transmembrane protein, across all stages of EOC. Expression was confirmed at both the mRNA and protein levels using RT-PCR and immunohistochemical staining, respectively.
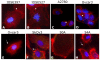
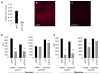
Results: In both EOC tumors and normal ovarian surface epithelial cells we demonstrated CD151 to be localized to the membrane and cell-cell junctions in patient-derived and established EOC cell lines. We next evaluated its role in EOC dissemination using two ovarian cancer-derived cell lines with differential levels of CD151 expression. Targeted antibody-mediated and siRNA inhibition or loss of CD151 in SKOV3 and OVCAR5 cell lines effectively inhibited their migration and invasion.
Conclusion: Taken together, these findings provide the first proof-of-principle demonstration for a next generation sequencing approach to identifying candidate therapeutic targets and reveal CD151 to play a role in EOC dissemination.
Figures

References
-
- Altekruse SF, Kosary CL, Krapcho M, Neyman N, Aminou R, Waldron W, Ruhl J, Howlader N, Tatalovich Z, Cho H, Mariotto A, Eisner MP, Lewis DR, Cronin K, Chen HS, Feuer EJ, Stinchcomb DG, Edwards BK. SEER Cancer Statistics Review, 1975-2007 [Internet] Bethesda, MD: National Cancer Institute;
-
- Pleasance ED, Stephens PJ, O'Meara S, McBride DJ, Meynert A, Jones D, Lin ML, Beare D, Lau KW, Greenman C, Varela I, Nik-Zainal S, Davies HR, Ordonez GR, Mudie LJ, Latimer C, Edkins S, Stebbings L, Chen L, Jia M, Leroy C, Marshall J, Menzies A, Butler A, Teague JW, Mangion J, Sun YA, McLaughlin SF, Peckham HE, Tsung EF, Costa GL, Lee CC, Minna JD, Gazdar A, Birney E, Rhodes MD, McKernan KJ, Stratton MR, Futreal PA, Campbell PJ. A small-cell lung cancer genome with complex signatures of tobacco exposure. Nature. 2010;463:184–90. doi: 10.1038/nature08629. - DOI - PMC - PubMed
-
- Timmermann B, Kerick M, Roehr C, Fischer A, Isau M, Boerno ST, Wunderlich A, Barmeyer C, Seemann P, Koenig J, Lappe M, Kuss AW, Garshasbi M, Bertram L, Trappe K, Werber M, Herrmann BG, Zatloukal K, Lehrach H, Schweiger MR. Somatic mutation profiles of MSI and MSS colorectal cancer identified by whole exome next generation sequencing and bioinformatics analysis. PLoS One. 2010;5:e15661. doi: 10.1371/journal.pone.0015661. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

