Progressive polycomb assembly on H3K27me3 compartments generates polycomb bodies with developmentally regulated motion
- PMID: 22275876
- PMCID: PMC3262012
- DOI: 10.1371/journal.pgen.1002465
Progressive polycomb assembly on H3K27me3 compartments generates polycomb bodies with developmentally regulated motion
Abstract
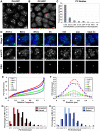
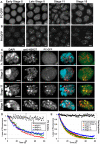
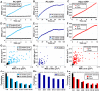
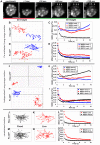
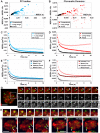
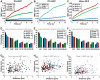
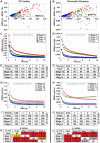
Polycomb group (PcG) proteins are conserved chromatin factors that maintain silencing of key developmental genes outside of their expression domains. Recent genome-wide analyses showed a Polycomb (PC) distribution with binding to discrete PcG response elements (PREs). Within the cell nucleus, PcG proteins localize in structures called PC bodies that contain PcG-silenced genes, and it has been recently shown that PREs form local and long-range spatial networks. Here, we studied the nuclear distribution of two PcG proteins, PC and Polyhomeotic (PH). Thanks to a combination of immunostaining, immuno-FISH, and live imaging of GFP fusion proteins, we could analyze the formation and the mobility of PC bodies during fly embryogenesis as well as compare their behavior to that of the condensed fraction of euchromatin. Immuno-FISH experiments show that PC bodies mainly correspond to 3D structural counterparts of the linear genomic domains identified in genome-wide studies. During early embryogenesis, PC and PH progressively accumulate within PC bodies, which form nuclear structures localized on distinct euchromatin domains containing histone H3 tri-methylated on K27. Time-lapse analysis indicates that two types of motion influence the displacement of PC bodies and chromatin domains containing H2Av-GFP. First, chromatin domains and PC bodies coordinately undergo long-range motions that may correspond to the movement of whole chromosome territories. Second, each PC body and chromatin domain has its own fast and highly constrained motion. In this motion regime, PC bodies move within volumes slightly larger than those of condensed chromatin domains. Moreover, both types of domains move within volumes much smaller than chromosome territories, strongly restricting their possibility of interaction with other nuclear structures. The fast motion of PC bodies and chromatin domains observed during early embryogenesis strongly decreases in late developmental stages, indicating a possible contribution of chromatin dynamics in the maintenance of stable gene silencing.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293:1074–1080. - PubMed
-
- Shao Z, Raible F, Mollaaghababa R, Guyon JR, Wu CT, et al. Stabilization of chromatin structure by PRC1, a Polycomb complex. Cell. 1999;98:37–46. - PubMed
-
- Simon JA, Kingston RE. Mechanisms of Polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol 2009 - PubMed
-
- Schwartz YB, Pirrotta V. Polycomb silencing mechanisms and the management of genomic programmes. Nat Rev Genet. 2007;8:9–22. - PubMed
-
- Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G. Genome regulation by polycomb and trithorax proteins. Cell. 2007;128:735–745. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

