Manipulating protein conformations by single-molecule AFM-FRET nanoscopy
- PMID: 22276737
- PMCID: PMC3662055
- DOI: 10.1021/nn2038669
Manipulating protein conformations by single-molecule AFM-FRET nanoscopy
Abstract
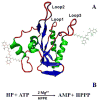
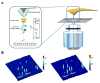
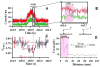
Combining atomic force microscopy and fluorescence resonance energy transfer spectroscopy (AFM-FRET), we have developed a single-molecule AFM-FRET nanoscopy approach capable of effectively pinpointing and mechanically manipulating a targeted dye-labeled single protein in a large sampling area and simultaneously monitoring the conformational changes of the targeted protein by recording single-molecule FRET time trajectories. We have further demonstrated an application of using this nanoscopy on manipulation of single-molecule protein conformation and simultaneous single-molecule FRET measurement of a Cy3-Cy5-labeled kinase enzyme, HPPK (6-hydroxymethyl-7,8-dihydropterin pyrophosphokinase). By analyzing time-resolved FRET trajectories and correlated AFM force pulling curves of the targeted single-molecule enzyme, we are able to observe the protein conformational changes of a specific coordination by AFM mechanic force pulling.
Figures
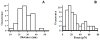

Similar articles
-
Manipulating and probing enzymatic conformational fluctuations and enzyme-substrate interactions by single-molecule FRET-magnetic tweezers microscopy.Phys Chem Chem Phys. 2014 Jul 14;16(26):13052-8. doi: 10.1039/c4cp01454e. Phys Chem Chem Phys. 2014. PMID: 24853252
-
Revealing Abrupt and Spontaneous Ruptures of Protein Native Structure under picoNewton Compressive Force Manipulation.ACS Nano. 2018 Mar 27;12(3):2448-2454. doi: 10.1021/acsnano.7b07934. Epub 2018 Mar 2. ACS Nano. 2018. PMID: 29462552
-
Single-molecule photon stamping FRET spectroscopy study of enzymatic conformational dynamics.Phys Chem Chem Phys. 2013 Jan 21;15(3):770-5. doi: 10.1039/c2cp42944f. Phys Chem Chem Phys. 2013. PMID: 23085845 Free PMC article.
-
Investigating piconewton forces in cells by FRET-based molecular force microscopy.J Struct Biol. 2017 Jan;197(1):37-42. doi: 10.1016/j.jsb.2016.03.011. Epub 2016 Mar 12. J Struct Biol. 2017. PMID: 26980477 Review.
-
Frontiers in microbial nanoscopy.Nanomedicine (Lond). 2011 Feb;6(2):395-403. doi: 10.2217/nnm.10.151. Nanomedicine (Lond). 2011. PMID: 21385140 Review.
Cited by
-
Method for high frequency tracking and sub-nm sample stabilization in single molecule fluorescence microscopy.Sci Rep. 2018 Sep 17;8(1):13912. doi: 10.1038/s41598-018-32012-1. Sci Rep. 2018. PMID: 30224660 Free PMC article.
-
Quantifying molecular- to cellular-level forces in living cells.J Phys D Appl Phys. 2021 Dec 2;54(48):483001. doi: 10.1088/1361-6463/ac2170. Epub 2021 Sep 9. J Phys D Appl Phys. 2021. PMID: 34866655 Free PMC article.
-
Self-Healing Ability of Perovskites Observed via Photoluminescence Response on Nanoscale Local Forces and Mechanical Damage.Adv Sci (Weinh). 2022 Dec 1;10(1):e2204393. doi: 10.1002/advs.202204393. Online ahead of print. Adv Sci (Weinh). 2022. PMID: 36453591 Free PMC article.
-
Phosphorylation Induces Conformational Rigidity at the C-Terminal Domain of AMPA Receptors.J Phys Chem B. 2019 Jan 10;123(1):130-137. doi: 10.1021/acs.jpcb.8b10749. Epub 2018 Dec 27. J Phys Chem B. 2019. PMID: 30537817 Free PMC article.
-
Single-molecule studies of riboswitch folding.Biochim Biophys Acta. 2014 Oct;1839(10):1030-1045. doi: 10.1016/j.bbagrm.2014.04.005. Epub 2014 Apr 13. Biochim Biophys Acta. 2014. PMID: 24727093 Free PMC article. Review.
References
-
- Boehr DD, Dyson HJ, Wright PE. An NMR Perspective on Enzyme Dynamics. Chem Rev. 2006;106:3055–3079. - PubMed
-
- Henzler-Wildman KA, Lei M, Thai V, Kerns SJ, Karplus M, Kern D. A Hierarchy of Timescales in Protein Dynamics Is Linked to Enzyme Catalysis. Nature. 2007;450:913–916. - PubMed
-
- Henzler-Wildman KA, Thai V, Lei M, Ott M, Wolf-Watz M, Fenn T, Pozharski E, Wilson MA, Petsko GA, Karplus M, Hubner CG, Kern D. Intrinsic Motions Along an Enzymatic Reaction Trajectory. Nature. 2007;450:838–844. - PubMed
-
- Min W, English BP, Luo GB, Cherayil BJ, Kou SC, Xie XS. Fluctuating Enzymes: Lessons from Single-Molecule Studies. Acc Chem Res. 2005;38:923–931. - PubMed
-
- Blaszczyk J, Li Y, Wu Y, Shi GB, Ji XH, Yan HG. Essential Roles of a Dynamic Loop in the Catalysis of 6-Hydroxymethyl-7,8-Dihydropterin Pyrophosphokinase. Biochemistry. 2004;43:1469–1477. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

