A genome triplication associated with early diversification of the core eudicots
- PMID: 22280555
- PMCID: PMC3334584
- DOI: 10.1186/gb-2012-13-1-r3
A genome triplication associated with early diversification of the core eudicots
Abstract
Background: Although it is agreed that a major polyploidy event, gamma, occurred within the eudicots, the phylogenetic placement of the event remains unclear.
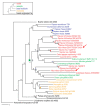
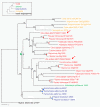
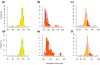
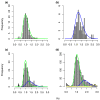
Results: To determine when this polyploidization occurred relative to speciation events in angiosperm history, we employed a phylogenomic approach to investigate the timing of gene set duplications located on syntenic gamma blocks. We populated 769 putative gene families with large sets of homologs obtained from public transcriptomes of basal angiosperms, magnoliids, asterids, and more than 91.8 gigabases of new next-generation transcriptome sequences of non-grass monocots and basal eudicots. The overwhelming majority (95%) of well-resolved gamma duplications was placed before the separation of rosids and asterids and after the split of monocots and eudicots, providing strong evidence that the gamma polyploidy event occurred early in eudicot evolution. Further, the majority of gene duplications was placed after the divergence of the Ranunculales and core eudicots, indicating that the gamma appears to be restricted to core eudicots. Molecular dating estimates indicate that the duplication events were intensely concentrated around 117 million years ago.
Conclusions: The rapid radiation of core eudicot lineages that gave rise to nearly 75% of angiosperm species appears to have occurred coincidentally or shortly following the gamma triplication event. Reconciliation of gene trees with a species phylogeny can elucidate the timing of major events in genome evolution, even when genome sequences are only available for a subset of species represented in the gene trees. Comprehensive transcriptome datasets are valuable complements to genome sequences for high-resolution phylogenomic analysis.
Figures

References
-
- Ohno S. Evolution by Gene Duplication. Springer-Verlag; 1970.
-
- Aury JM, Jaillon O, Duret L, Noel B, Jubin C, Porcel BM, Segurens B, Daubin V, Anthouard V, Aiach N, Arnaiz O, Billaut A, Beisson J, Blanc I, Bouhouche K, Camara F, Duharcourt S, Guigo R, Gogendeau D, Katinka M, Keller AM, Kissmehl R, Klotz C, Koll F, Le Mouel A, Lepere G, Malinsky S, Nowacki M, Nowak JK, Plattner H. et al. Global trends of whole-genome duplications revealed by the ciliate Paramecium tetraurelia. Nature. 2006;444:171–178. doi: 10.1038/nature05230. - DOI - PubMed
-
- Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, dePamphilis CW. Ancestral polyploidy in seed plants and angiosperms. Nature. 2011;473:97–100. doi: 10.1038/nature09916. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

