Multiple roles of class I HDACs in proliferation, differentiation, and development
- PMID: 22286122
- PMCID: PMC11115120
- DOI: 10.1007/s00018-012-0921-9
Multiple roles of class I HDACs in proliferation, differentiation, and development
Abstract
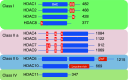
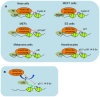
Class I Histone deacetylases (HDACs) play a central role in controlling cell cycle regulation, cell differentiation, and tissue development. These enzymes exert their function by deacetylating histones and a growing number of non-histone proteins, thereby regulating gene expression and several other cellular processes. Class I HDACs comprise four members: HDAC1, 2, 3, and 8. Deletion and/or overexpression of these enzymes in mammalian systems has provided important insights about their functions and mechanisms of action which are reviewed here. In particular, unique as well as redundant functions have been identified in several paradigms. Studies with small molecule inhibitors of HDACs have demonstrated the medical relevance of these enzymes and their potential as therapeutic targets in cancer and other pathological conditions. Going forward, better understanding the specific role of individual HDACs in normal physiology as well as in pathological settings will be crucial to exploit this protein family as a useful therapeutic target in a range of diseases. Further dissection of the pathways they impinge on and of their targets, in chromatin or otherwise, will form important avenues of research for the future.
Figures

Similar articles
-
Targeting Class I Histone Deacetylases in Human Uterine Leiomyosarcoma.Cells. 2022 Nov 27;11(23):3801. doi: 10.3390/cells11233801. Cells. 2022. PMID: 36497061 Free PMC article.
-
Targeting histone deacetylases for cancer therapy: from molecular mechanisms to clinical implications.Int J Biol Sci. 2014 Jul 2;10(7):757-70. doi: 10.7150/ijbs.9067. eCollection 2014. Int J Biol Sci. 2014. PMID: 25013383 Free PMC article. Review.
-
Histone deacetylase (HDAC) activity is critical for embryonic kidney gene expression, growth, and differentiation.J Biol Chem. 2011 Sep 16;286(37):32775-89. doi: 10.1074/jbc.M111.248278. Epub 2011 Jul 21. J Biol Chem. 2011. PMID: 21778236 Free PMC article.
-
Histone deacetylases as targets for treatment of multiple diseases.Clin Sci (Lond). 2013 Jun;124(11):651-62. doi: 10.1042/CS20120504. Clin Sci (Lond). 2013. PMID: 23414309 Free PMC article. Review.
-
Protein deacetylases: enzymes with functional diversity as novel therapeutic targets.Prog Cell Cycle Res. 2003;5:269-78. Prog Cell Cycle Res. 2003. PMID: 14593721 Review.
Cited by
-
Prefrontal cortical dysfunction after overexpression of histone deacetylase 1.Biol Psychiatry. 2013 Nov 1;74(9):696-705. doi: 10.1016/j.biopsych.2013.03.020. Epub 2013 May 7. Biol Psychiatry. 2013. PMID: 23664640 Free PMC article.
-
Targeting Class I Histone Deacetylases in Human Uterine Leiomyosarcoma.Cells. 2022 Nov 27;11(23):3801. doi: 10.3390/cells11233801. Cells. 2022. PMID: 36497061 Free PMC article.
-
Histone Modifications and Non-Coding RNAs: Mutual Epigenetic Regulation and Role in Pathogenesis.Int J Mol Sci. 2022 May 22;23(10):5801. doi: 10.3390/ijms23105801. Int J Mol Sci. 2022. PMID: 35628612 Free PMC article. Review.
-
Beyond Gut Instinct: Metabolic Short-Chain Fatty Acids Moderate the Pathogenesis of Alphaherpesviruses.Front Microbiol. 2019 Apr 5;10:723. doi: 10.3389/fmicb.2019.00723. eCollection 2019. Front Microbiol. 2019. PMID: 31024501 Free PMC article.
-
MicroRNAs: potential regulators of renal development genes that contribute to CAKUT.Pediatr Nephrol. 2014 Apr;29(4):565-74. doi: 10.1007/s00467-013-2599-0. Epub 2013 Sep 3. Pediatr Nephrol. 2014. PMID: 23996519 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

