The SAR11 group of alpha-proteobacteria is not related to the origin of mitochondria
- PMID: 22291975
- PMCID: PMC3264578
- DOI: 10.1371/journal.pone.0030520
The SAR11 group of alpha-proteobacteria is not related to the origin of mitochondria
Abstract
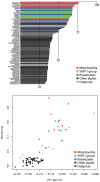
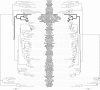
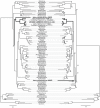
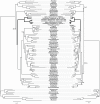
Although free living, members of the successful SAR11 group of marine alpha-proteobacteria contain a very small and A+T rich genome, two features that are typical of mitochondria and related obligate intracellular parasites such as the Rickettsiales. Previous phylogenetic analyses have suggested that Candidatus Pelagibacter ubique, the first cultured member of this group, is related to the Rickettsiales+mitochondria clade whereas others disagree with this conclusion. In order to determine the evolutionary position of the SAR11 group and its relationship to the origin of mitochondria, we have performed phylogenetic analyses on the concatenation of 24 proteins from 5 mitochondria and 71 proteobacteria. Our results support that SAR11 group is not the sistergroup of the Rickettsiales+mitochondria clade and confirm that the position of this group in the alpha-proteobacterial tree is strongly affected by tree reconstruction artefacts due to compositional bias. As a consequence, genome reduction and bias toward a high A+T content may have evolved independently in the SAR11 species, which points to a different direction in the quest for the closest relatives to mitochondria and Rickettsiales. In addition, our analyses raise doubts about the monophyly of the newly proposed Pelagibacteraceae family.
Conflict of interest statement
Figures
References
-
- Andersson SG, Zomorodipour A, Andersson JO, Sicheritz-Ponten T, Alsmark UC, et al. The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature. 1998;396:133–140. - PubMed
-
- Lang BF, Gray MW, Burger G. Mitochondrial genome evolution and the origin of eukaryotes. Annu Rev Genet. 1999;33:351–397. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

