Heparan sulfate sulfatase SULF2 regulates PDGFRα signaling and growth in human and mouse malignant glioma
- PMID: 22293178
- PMCID: PMC3287216
- DOI: 10.1172/JCI58215
Heparan sulfate sulfatase SULF2 regulates PDGFRα signaling and growth in human and mouse malignant glioma
Abstract
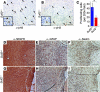
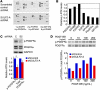
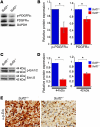
Glioblastoma (GBM), a uniformly lethal brain cancer, is characterized by diffuse invasion and abnormal activation of multiple receptor tyrosine kinase (RTK) signaling pathways, presenting a major challenge to effective therapy. The activation of many RTK pathways is regulated by extracellular heparan sulfate proteoglycans (HSPG), suggesting these molecules may be effective targets in the tumor microenvironment. In this study, we demonstrated that the extracellular sulfatase, SULF2, an enzyme that regulates multiple HSPG-dependent RTK signaling pathways, was expressed in primary human GBM tumors and cell lines. Knockdown of SULF2 in human GBM cell lines and generation of gliomas from Sulf2(-/-) tumorigenic neurospheres resulted in decreased growth in vivo in mice. We found a striking SULF2 dependence in activity of PDGFRα, a major signaling pathway in GBM. Ablation of SULF2 resulted in decreased PDGFRα phosphorylation and decreased downstream MAPK signaling activity. Interestingly, in a survey of SULF2 levels in different subtypes of GBM, the proneural subtype, characterized by aberrations in PDGFRα, demonstrated the strongest SULF2 expression. Therefore, in addition to its potential as an upstream target for therapy of GBM, SULF2 may help identify a subset of GBMs that are more dependent on exogenous growth factor-mediated signaling. Our results suggest the bioavailability of growth factors from the microenvironment is a significant contributor to tumor growth in a major subset of human GBM.
Figures






References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases

