Protein-binding dynamics imaged in a living cell
- PMID: 22307600
- PMCID: PMC3277120
- DOI: 10.1073/pnas.1112171109
Protein-binding dynamics imaged in a living cell
Abstract
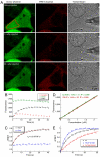
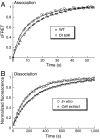
Historically, rate constants were determined in vitro and it was unknown whether they were valid for in vivo biological processes. Here, we bridge this gap by measuring binding dynamics between a pair of proteins in living HeLa cells. Binding of a β-lactamase to its protein inhibitor was initiated by microinjection and monitored by Förster resonance energy transfer. Association rate constants for the wild-type and an electrostatically optimized mutant were only 25% and 50% lower than in vitro values, whereas no change in the rate constant was observed for a slower binding mutant. These changes are much smaller than might be anticipated considering the high macromolecular crowding within the cell. Single-cell analyses of association rate constants and fluorescence recovery after photobleaching reveals a naturally occurring variation in cell density, which is translated to an up to a twofold effect on binding rate constants. The data show that for this model protein interaction the intracellular environment had only a small effect on the association kinetics, justifying the extrapolation of in vitro data to processes in the cell.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

