Structural reorganization of the interleukin-7 signaling complex
- PMID: 22308406
- PMCID: PMC3289338
- DOI: 10.1073/pnas.1116582109
Structural reorganization of the interleukin-7 signaling complex
Abstract
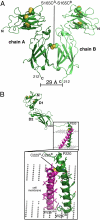
We report here an unliganded receptor structure in the common gamma-chain (γ(c)) family of receptors and cytokines. The crystal structure of the unliganded form of the interleukin-7 alpha receptor (IL-7Rα) extracellular domain (ECD) at 2.15 Å resolution reveals a homodimer forming an "X" geometry looking down onto the cell surface with the C termini of the two chains separated by 110 Å and the dimer interface comprising residues critical for IL-7 binding. Further biophysical studies indicate a weak association of the IL-7Rα ECDs but a stronger association between the γ(c)/IL-7Rα ECDs, similar to previous studies of the full-length receptors on CD4(+) T cells. Based on these and previous results, we propose a molecular mechanism detailing the progression from the inactive IL-7Rα homodimer and IL-7Rα-γ(c) heterodimer to the active IL-7-IL-7Rα-γ(c) ternary complex whereby the two receptors undergo at least a 90° rotation away from the cell surface, moving the C termini of IL-7Rα and γ(c) from a distance of 110 Å to less than 30 Å at the cell surface. This molecular mechanism can be used to explain recently discovered IL-7- and γ(c)-independent gain-of-function mutations in IL-7Rα from B- and T-cell acute lymphoblastic leukemia patients. The mechanism may also be applicable to other γ(c) receptors that form inactive homodimers and heterodimers independent of their cytokines.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Structural insights into the common γ-chain family of cytokines and receptors from the interleukin-7 pathway.Immunol Rev. 2012 Nov;250(1):303-16. doi: 10.1111/j.1600-065X.2012.01160.x. Immunol Rev. 2012. PMID: 23046137 Free PMC article. Review.
-
Structural and biophysical studies of the human IL-7/IL-7Ralpha complex.Structure. 2009 Jan 14;17(1):54-65. doi: 10.1016/j.str.2008.10.019. Structure. 2009. PMID: 19141282 Free PMC article.
-
A biosensor study indicating that entropy, electrostatics, and receptor glycosylation drive the binding interaction between interleukin-7 and its receptor.Biochemistry. 2010 Oct 12;49(40):8766-78. doi: 10.1021/bi101050h. Epub 2010 Sep 15. Biochemistry. 2010. PMID: 20815339 Free PMC article.
-
IL-7 Induces an Epitope Masking of γc Protein in IL-7 Receptor Signaling Complex.Mediators Inflamm. 2017;2017:9096829. doi: 10.1155/2017/9096829. Epub 2017 Jan 3. Mediators Inflamm. 2017. PMID: 28127156 Free PMC article.
-
Therapeutic targeting of IL-7Rα signaling pathways in ALL treatment.Blood. 2016 Jul 28;128(4):473-8. doi: 10.1182/blood-2016-03-679209. Epub 2016 Jun 6. Blood. 2016. PMID: 27268088 Free PMC article. Review.
Cited by
-
Crystal structure of Lamellipodin implicates diverse functions in actin polymerization and Ras signaling.Protein Cell. 2013 Mar;4(3):211-9. doi: 10.1007/s13238-013-2082-5. Epub 2013 Mar 13. Protein Cell. 2013. PMID: 23483482 Free PMC article.
-
Membrane microdomains and cytoskeleton organization shape and regulate the IL-7 receptor signalosome in human CD4 T-cells.J Biol Chem. 2013 Mar 22;288(12):8691-8701. doi: 10.1074/jbc.M113.449918. Epub 2013 Jan 17. J Biol Chem. 2013. PMID: 23329834 Free PMC article.
-
The Cytokine Receptor IL-7Rα Impairs IL-2 Receptor Signaling and Constrains the In Vitro Differentiation of Foxp3+ Treg Cells.iScience. 2020 Aug 21;23(8):101421. doi: 10.1016/j.isci.2020.101421. Epub 2020 Jul 29. iScience. 2020. PMID: 32791329 Free PMC article.
-
Association of interleukin 7 receptor gene polymorphism rs6897932 with multiple sclerosis patients in Khuzestan.Iran J Neurol. 2014 Jul 4;13(3):168-71. Iran J Neurol. 2014. PMID: 25422737 Free PMC article.
-
The common γ-chain cytokine receptor: tricks-and-treats for T cells.Cell Mol Life Sci. 2016 Jan;73(2):253-69. doi: 10.1007/s00018-015-2062-4. Epub 2015 Oct 14. Cell Mol Life Sci. 2016. PMID: 26468051 Free PMC article. Review.
References
-
- Mazzucchelli R, Durum SK. Interleukin-7 receptor expression: Intelligent design. Nat Rev Immunol. 2007;7(2):144–154. - PubMed
-
- Ozaki K, Leonard WJ. Cytokine and cytokine receptor pleiotropy and redundancy. J Biol Chem. 2002;277:29355–29358. - PubMed
-
- Kovanen PE, Leonard WJ. Cytokines and immunodeficiency diseases: Critical roles of the γ(c)-dependent cytokines interleukins 2, 4, 7, 9, 15, and 21, and their signaling pathways. Immunol Rev. 2004;202(1):67–83. - PubMed
-
- Gregory SG, et al. Multiple Sclerosis Genetics Group. Interleukin 7 receptor α chain (IL7R) shows allelic and functional association with multiple sclerosis. Nat Genet. 2007;39:1083–1091. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

