Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types
- PMID: 22319583
- PMCID: PMC3270023
- DOI: 10.1371/journal.pone.0030733
Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types
Abstract
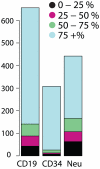
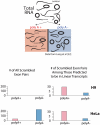
Most human pre-mRNAs are spliced into linear molecules that retain the exon order defined by the genomic sequence. By deep sequencing of RNA from a variety of normal and malignant human cells, we found RNA transcripts from many human genes in which the exons were arranged in a non-canonical order. Statistical estimates and biochemical assays provided strong evidence that a substantial fraction of the spliced transcripts from hundreds of genes are circular RNAs. Our results suggest that a non-canonical mode of RNA splicing, resulting in a circular RNA isoform, is a general feature of the gene expression program in human cells.
Conflict of interest statement
Figures




References
-
- Coca-Prados M-THaM. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979;280:339–340. - PubMed
-
- Nigro JM, Cho KR, Fearon ER, Kern SE, Ruppert JM, et al. Scrambled exons. Cell. 1991;64:607–613. - PubMed
-
- Saad FA, Vitiello L, Merlini L, Mostacciuolo ML, Oliviero S, et al. A 3′ consensus splice mutation in the human dystrophin gene detected by a screening for intra-exonic deletions. Human molecular genetics. 1992;1:345–346. - PubMed
-
- Capel B, Swain A, Nicolis S, Hacker A, Walter M, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–1030. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

