iSyTE: integrated Systems Tool for Eye gene discovery
- PMID: 22323457
- PMCID: PMC3339920
- DOI: 10.1167/iovs.11-8839
iSyTE: integrated Systems Tool for Eye gene discovery
Abstract
Purpose: To facilitate the identification of genes associated with cataract and other ocular defects, the authors developed and validated a computational tool termed iSyTE (integrated Systems Tool for Eye gene discovery; http://bioinformatics.udel.edu/Research/iSyTE). iSyTE uses a mouse embryonic lens gene expression data set as a bioinformatics filter to select candidate genes from human or mouse genomic regions implicated in disease and to prioritize them for further mutational and functional analyses.
Methods: Microarray gene expression profiles were obtained for microdissected embryonic mouse lens at three key developmental time points in the transition from the embryonic day (E)10.5 stage of lens placode invagination to E12.5 lens primary fiber cell differentiation. Differentially regulated genes were identified by in silico comparison of lens gene expression profiles with those of whole embryo body (WB) lacking ocular tissue.
Results: Gene set analysis demonstrated that this strategy effectively removes highly expressed but nonspecific housekeeping genes from lens tissue expression profiles, allowing identification of less highly expressed lens disease-associated genes. Among 24 previously mapped human genomic intervals containing genes associated with isolated congenital cataract, the mutant gene is ranked within the top two iSyTE-selected candidates in approximately 88% of cases. Finally, in situ hybridization confirmed lens expression of several novel iSyTE-identified genes.
Conclusions: iSyTE is a publicly available Web resource that can be used to prioritize candidate genes within mapped genomic intervals associated with congenital cataract for further investigation. Extension of this approach to other ocular tissue components will facilitate eye disease gene discovery.
Figures
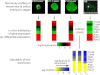




References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

