Co-transcriptional regulation of alternative pre-mRNA splicing
- PMID: 22326677
- PMCID: PMC3371144
- DOI: 10.1016/j.bbagrm.2012.01.014
Co-transcriptional regulation of alternative pre-mRNA splicing
Abstract
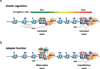
While studies of alternative pre-mRNA splicing regulation have typically focused on RNA-binding proteins and their target sequences within nascent message, it is becoming increasingly evident that mRNA splicing, RNA polymerase II (pol II) elongation and chromatin structure are intricately intertwined. The majority of introns in higher eukaryotes are excised prior to transcript release in a manner that is dependent on transcription through pol II. As a result of co-transcriptional splicing, variations in pol II elongation influence alternative splicing patterns, wherein a slower elongation rate is associated with increased inclusion of alternative exons within mature mRNA. Physiological barriers to pol II elongation, such as repressive chromatin structure, can thereby similarly impact splicing decisions. Surprisingly, pre-mRNA splicing can reciprocally influence pol II elongation and chromatin structure. Here, we highlight recent advances in co-transcriptional splicing that reveal an extensive network of coupling between splicing, transcription and chromatin remodeling complexes. This article is part of a Special Issue entitled: Chromatin in time and space.
Published by Elsevier B.V.
Figures


Similar articles
-
Differential patterns of intronic and exonic DNA regions with respect to RNA polymerase II occupancy, nucleosome density and H3K36me3 marking in fission yeast.Genome Biol. 2011 Aug 22;12(8):R82. doi: 10.1186/gb-2011-12-8-r82. Genome Biol. 2011. PMID: 21859475 Free PMC article.
-
On the importance of being co-transcriptional.J Cell Sci. 2002 Oct 15;115(Pt 20):3865-71. doi: 10.1242/jcs.00073. J Cell Sci. 2002. PMID: 12244124 Review.
-
The in vivo kinetics of RNA polymerase II elongation during co-transcriptional splicing.PLoS Biol. 2011 Jan 11;9(1):e1000573. doi: 10.1371/journal.pbio.1000573. PLoS Biol. 2011. PMID: 21264352 Free PMC article.
-
Coupling of RNA Polymerase II Transcription Elongation with Pre-mRNA Splicing.J Mol Biol. 2016 Jun 19;428(12):2623-2635. doi: 10.1016/j.jmb.2016.04.017. Epub 2016 Apr 20. J Mol Biol. 2016. PMID: 27107644 Free PMC article. Review.
-
Coupling between transcription and alternative splicing.Cancer Treat Res. 2013;158:1-24. doi: 10.1007/978-3-642-31659-3_1. Cancer Treat Res. 2013. PMID: 24222352
Cited by
-
Inorganic Arsenic-induced cellular transformation is coupled with genome wide changes in chromatin structure, transcriptome and splicing patterns.BMC Genomics. 2015 Mar 19;16(1):212. doi: 10.1186/s12864-015-1295-9. BMC Genomics. 2015. PMID: 25879800 Free PMC article.
-
Dietary-phytochemical mediated reversion of cancer-specific splicing inhibits Warburg effect in head and neck cancer.BMC Cancer. 2019 Nov 1;19(1):1031. doi: 10.1186/s12885-019-6257-1. BMC Cancer. 2019. PMID: 31675998 Free PMC article.
-
Beyond Transcription: Roles of Transcription Factors in Pre-mRNA Splicing.Chem Rev. 2018 Apr 25;118(8):4339-4364. doi: 10.1021/acs.chemrev.7b00470. Epub 2017 Dec 18. Chem Rev. 2018. PMID: 29251915 Free PMC article. Review.
-
Chromatin modifier MTA1 regulates mitotic transition and tumorigenesis by orchestrating mitotic mRNA processing.Nat Commun. 2020 Sep 8;11(1):4455. doi: 10.1038/s41467-020-18259-1. Nat Commun. 2020. PMID: 32901005 Free PMC article.
-
Intron retention in the alternatively spliced region of RON results from weak 3' splice site recognition.PLoS One. 2013 Oct 14;8(10):e77208. doi: 10.1371/journal.pone.0077208. eCollection 2013. PLoS One. 2013. PMID: 24155930 Free PMC article.
References
-
- Chow LT, Gelinas RE, Broker TR, Roberts RJ. An amazing sequence arrangement at the 5' ends of adenovirus 2 messenger RNA. Cell. 1977;12:1–8. - PubMed
-
- Pan Q, Shai O, Lee LJ, Frey BJ, Blencowe BJ. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat Genet. 2008;40:1413–1415. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

