Dual transcriptional activator and repressor roles of TBX20 regulate adult cardiac structure and function
- PMID: 22328084
- PMCID: PMC3335310
- DOI: 10.1093/hmg/dds034
Dual transcriptional activator and repressor roles of TBX20 regulate adult cardiac structure and function
Abstract
The ongoing requirement in adult heart for transcription factors with key roles in cardiac development is not well understood. We recently demonstrated that TBX20, a transcriptional regulator required for cardiac development, has key roles in the maintenance of functional and structural phenotypes in adult mouse heart. Conditional ablation of Tbx20 in adult cardiomyocytes leads to a rapid onset and progression of heart failure, with prominent conduction and contractility phenotypes that lead to death. Here we describe a more comprehensive molecular characterization of the functions of TBX20 in adult mouse heart. Coupling genome-wide chromatin immunoprecipitation and transcriptome analyses (RNA-Seq), we identified a subset of genes that change expression in Tbx20 adult cardiomyocyte-specific knockout hearts which are direct downstream targets of TBX20. This analysis revealed a dual role for TBX20 as both a transcriptional activator and a repressor, and that each of these functions regulates genes with very specialized and distinct molecular roles. We also show how TBX20 binds to its targets genome-wide in a context-dependent manner, using various cohorts of co-factors to either promote or repress distinct genetic programs within adult heart. Our integrative approach has uncovered several novel aspects of TBX20 and T-box protein function within adult heart. Sequencing data accession number (http://www.ncbi.nlm.nih.gov/geo): GSE30943.
Figures

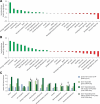



References
-
- Singh M.K., Christoffels V.M., Dias J.M., Trowe M.-O., Petry M., Schuster-Gossler K., Bürger A., Ericson J., Kispert A. Tbx20 is essential for cardiac chamber differentiation and repression of Tbx2. Development. 2005;132:2697–2707. - PubMed
-
- Stennard F.A., Costa M.W., Lai D., Biben C., Furtado M.B., Solloway M.J., McCulley D.J., Leimena C., Preis J.I., Dunwoodie S.L., et al. Murine T-box transcription factor Tbx20 acts as a repressor during heart development, and is essential for adult heart integrity, function and adaptation. Development. 2005;132:2451–2462. - PubMed
-
- Takeuchi J.K., Mileikovskaia M., Koshiba-Takeuchi K., Heidt A.B., Mori A.D., Arruda E.P., Gertsenstein M., Georges R., Davidson L., Mo R., et al. Tbx20 dose-dependently regulates transcription factor networks required for mouse heart and motoneuron development. Development. 2005;132:2463–2474. - PubMed
-
- Liu C., Shen A., Li X., Jiao W., Zhang X., Li Z. T-box transcription factor TBX20 mutations in Chinese patients with congenital heart disease. Eur. J. Med. Genet. 2008;51:580–587. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

