Difference in fine specificity to polysaccharides of Candida albicans mannoprotein between mouse SIGNR1 and human DC-SIGN
- PMID: 22331432
- PMCID: PMC3347427
- DOI: 10.1128/IAI.06308-11
Difference in fine specificity to polysaccharides of Candida albicans mannoprotein between mouse SIGNR1 and human DC-SIGN
Abstract
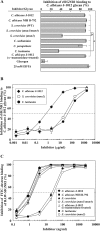
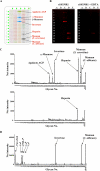
C-type lectin SIGNR1 directly recognizes Candida albicans and zymosan and has been considered to share properties of polysaccharide recognition with human DC-SIGN (hDC-SIGN). However, the precise specificity of SIGNR1 and the difference from that of hDC-SIGN remain to be elucidated. We prepared soluble forms of SIGNR1 and hDC-SIGN and conducted experiments to examine their respective specificities. Soluble SIGNR1 (sSIGNR1) bound several types of live C. albicans clinical isolate strains in an EDTA-sensitive manner. Inhibition analyses of sSIGNR1 binding by glycans from various yeast strains demonstrated that SIGNR1 preferentially recognizes N-glycan α-mannose side chains in Candida mannoproteins, as reported in hDC-SIGN. Unlike shDC-SIGN, however, sSIGNR1 recognized not only Saccharomyces cerevisiae, but also C. albicans J-1012 glycan, even after α-mannosidase treatment that leaves only β1,2-mannose-capped α-mannose side chains. In addition, glycomicroarray analyses showed that sSIGNR1 binds mannans from C. albicans and S. cerevisiae but does not recognize Lewis(a/b/x/y) antigen polysaccharides as in shDC-SIGN. Consistent with these results, RAW264.7 cells expressing hDC-SIGN in which the carbohydrate recognition domain (CRD) was replaced with that of SIGNR1 (RAW-chimera) produced comparable amounts of interleukin 10 (IL-10) in response to glycans from C. albicans and S. cerevisiae, but those expressing hDC-SIGN produced less IL-10 in response to S. cerevisiae than C. albicans. Furthermore, RAW-hDC-SIGN cells remarkably reduced IL-10 production after α-mannosidase treatment compared with RAW-chimera cells. These results indicate that SIGNR1 recognizes C. albicans/yeast through a specificity partly distinct from that of its homologue hDC-SIGN.
Figures



Similar articles
-
C-type lectin SIGNR1 enhances cellular oxidative burst response against C. albicans in cooperation with Dectin-1.Eur J Immunol. 2011 May;41(5):1435-44. doi: 10.1002/eji.200940188. Epub 2011 Mar 14. Eur J Immunol. 2011. PMID: 21400494
-
Efficient capture of Candida albicans and zymosan by SIGNR1 augments TLR2-dependent TNF-α production.Int Immunol. 2012 Feb;24(2):89-96. doi: 10.1093/intimm/dxr103. Epub 2011 Dec 29. Int Immunol. 2012. PMID: 22207132
-
The role of SIGNR1 and the beta-glucan receptor (dectin-1) in the nonopsonic recognition of yeast by specific macrophages.J Immunol. 2004 Jan 15;172(2):1157-62. doi: 10.4049/jimmunol.172.2.1157. J Immunol. 2004. PMID: 14707091
-
DC-SIGN and mannosylated surface structures of Mycobacterium tuberculosis: a deceptive liaison.Eur J Cell Biol. 2010 Jan;89(1):95-101. doi: 10.1016/j.ejcb.2009.10.004. Epub 2009 Nov 4. Eur J Cell Biol. 2010. PMID: 19892432 Review.
-
Glycan-based DC-SIGN targeting vaccines to enhance antigen cross-presentation.Mol Immunol. 2013 Sep;55(2):143-5. doi: 10.1016/j.molimm.2012.10.031. Epub 2012 Nov 14. Mol Immunol. 2013. PMID: 23158834 Review.
Cited by
-
The Interaction of Human Pathogenic Fungi With C-Type Lectin Receptors.Front Immunol. 2018 Jun 4;9:1261. doi: 10.3389/fimmu.2018.01261. eCollection 2018. Front Immunol. 2018. PMID: 29915598 Free PMC article. Review.
-
Innate Defense against Fungal Pathogens.Cold Spring Harb Perspect Med. 2014 Nov 10;5(6):a019620. doi: 10.1101/cshperspect.a019620. Cold Spring Harb Perspect Med. 2014. PMID: 25384766 Free PMC article. Review.
-
TETRALEC, Artificial Tetrameric Lectins: A Tool to Screen Ligand and Pathogen Interactions.Int J Mol Sci. 2020 Jul 25;21(15):5290. doi: 10.3390/ijms21155290. Int J Mol Sci. 2020. PMID: 32722514 Free PMC article.
-
Carbohydrate microarrays for screening functional glycans.Chem Sci. 2016 Mar 1;7(3):2084-2093. doi: 10.1039/c5sc03789a. Epub 2016 Jan 5. Chem Sci. 2016. PMID: 29899934 Free PMC article.
-
Cell wall N-glycan of Candida albicans ameliorates early hyper- and late hypo-immunoreactivity in sepsis.Commun Biol. 2021 Mar 16;4(1):342. doi: 10.1038/s42003-021-01870-3. Commun Biol. 2021. PMID: 33727664 Free PMC article.
References
-
- Ballou CE, Kern KA, Raschke WC. 1973. Genetic control of yeast mannan structure. Complementation studies and properties of mannan mutants. J. Biol. Chem. 248:4667–4671 - PubMed
-
- Dupasquier M, Stoitzner P, van Oudenaren A, Romani N, Leenen PJ. 2004. Macrophages and dendritic cells constitute a major subpopulation of cells in the mouse dermis. J. Investig. Dermatol. 123:876–879 - PubMed
-
- Galustian C, et al. 2004. High and low affinity carbohydrate ligands revealed for murine SIGN-R1 by carbohydrate array and cell binding approaches, and differing specificities for SIGN-R3 and langerin. Int. Immunol. 16:853–866 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

