Structural basis for specific recognition of Rpt1p, an ATPase subunit of 26 S proteasome, by proteasome-dedicated chaperone Hsm3p
- PMID: 22334676
- PMCID: PMC3320968
- DOI: 10.1074/jbc.M112.345876
Structural basis for specific recognition of Rpt1p, an ATPase subunit of 26 S proteasome, by proteasome-dedicated chaperone Hsm3p
Abstract
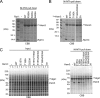
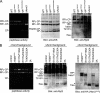
The 26 S proteasome is a 2.5-MDa molecular machine that degrades ubiquitinated proteins in eukaryotic cells. It consists of a proteolytic core particle and two 19 S regulatory particles (RPs) composed of 6 ATPase (Rpt) and 13 non-ATPase (Rpn) subunits. Multiple proteasome-dedicated chaperones facilitate the assembly of the proteasome, but little is known about the detailed mechanisms. Hsm3, a 19 S RP dedicated chaperone, transiently binds to the C-terminal domain of the Rpt1 subunit and forms a tetrameric complex, Hsm3-Rpt1-Rpt2-Rpn1, during maturation of the ATPase ring of 19 S RP. To elucidate the structural basis of Hsm3 function, we determined the crystal structures of Hsm3 and its complex with the C-terminal domain of the Rpt1 subunit (Rpt1C). Hsm3 has a C-shaped structure that consists of 11 HEAT repeats. The structure of the Hsm3-Rpt1C complex revealed that the interacting surface between Hsm3 and Rpt1 is a hydrophobic core and a complementary charged surface. Mutations in the Hsm3-Rpt1 surface resulted in the assembly defect of the 26 S proteasome. Furthermore, a structural model of the Hsm3-Rpt ring complex and an in vitro binding assay suggest that Hsm3 can bind Rpt2 in addition to Rpt1. Collectively, our results provide the structural basis of the molecular functions of Hsm3 for the RP assembly.
Figures




Similar articles
-
Dual functions of the Hsm3 protein in chaperoning and scaffolding regulatory particle subunits during the proteasome assembly.Proc Natl Acad Sci U S A. 2012 Apr 24;109(17):E1001-10. doi: 10.1073/pnas.1116538109. Epub 2012 Mar 29. Proc Natl Acad Sci U S A. 2012. PMID: 22460800 Free PMC article.
-
Hexameric assembly of the proteasomal ATPases is templated through their C termini.Nature. 2009 Jun 11;459(7248):866-70. doi: 10.1038/nature08065. Nature. 2009. PMID: 19412160 Free PMC article.
-
Crystal structure of yeast rpn14, a chaperone of the 19 S regulatory particle of the proteasome.J Biol Chem. 2010 May 14;285(20):15159-15166. doi: 10.1074/jbc.M110.104042. Epub 2010 Mar 16. J Biol Chem. 2010. PMID: 20236927 Free PMC article.
-
Assembly manual for the proteasome regulatory particle: the first draft.Biochem Soc Trans. 2010 Feb;38(Pt 1):6-13. doi: 10.1042/BST0380006. Biochem Soc Trans. 2010. PMID: 20074027 Free PMC article. Review.
-
Proteasome activator 200: the heat is on..Mol Cell Proteomics. 2011 May;10(5):R110.006890. doi: 10.1074/mcp.R110.006890. Epub 2011 Mar 9. Mol Cell Proteomics. 2011. PMID: 21389348 Free PMC article. Review.
Cited by
-
Proteasome gene expression is controlled by coordinated functions of multiple transcription factors.J Cell Biol. 2024 Aug 5;223(8):e202402046. doi: 10.1083/jcb.202402046. Epub 2024 May 20. J Cell Biol. 2024. PMID: 38767572 Free PMC article.
-
Dual functions of the Hsm3 protein in chaperoning and scaffolding regulatory particle subunits during the proteasome assembly.Proc Natl Acad Sci U S A. 2012 Apr 24;109(17):E1001-10. doi: 10.1073/pnas.1116538109. Epub 2012 Mar 29. Proc Natl Acad Sci U S A. 2012. PMID: 22460800 Free PMC article.
-
1.15 Å resolution structure of the proteasome-assembly chaperone Nas2 PDZ domain.Acta Crystallogr F Struct Biol Commun. 2014 Apr;70(Pt 4):418-23. doi: 10.1107/S2053230X14003884. Epub 2014 Mar 25. Acta Crystallogr F Struct Biol Commun. 2014. PMID: 24699731 Free PMC article.
-
Purification, crystallization and preliminary X-ray data collection of the N-terminal domain of the 26S proteasome regulatory subunit p27 and its complex with the ATPase domain of Rpt5 from Mus musculus.Acta Crystallogr F Struct Biol Commun. 2014 May;70(Pt 5):611-5. doi: 10.1107/S2053230X14006815. Epub 2014 Apr 15. Acta Crystallogr F Struct Biol Commun. 2014. PMID: 24817721 Free PMC article.
-
Structural insights on the dynamics of proteasome formation.Biophys Rev. 2018 Apr;10(2):597-604. doi: 10.1007/s12551-017-0381-4. Epub 2017 Dec 14. Biophys Rev. 2018. PMID: 29243089 Free PMC article. Review.
References
-
- Baumeister W., Walz J., Zühl F., Seemüller E. (1998) The proteasome. Paradigm of a self-compartmentalizing protease. Cell 92, 367–380 - PubMed
-
- Coux O., Tanaka K., Goldberg A. L. (1996) Structure and functions of the 20 S and 26 S proteasomes. Annu. Rev. Biochem. 65, 801–847 - PubMed
-
- Glickman M. H., Rubin D. M., Coux O., Wefes I., Pfeifer G., Cjeka Z., Baumeister W., Fried V. A., Finley D. (1998) A subcomplex of the proteasome regulatory particle required for ubiquitin-conjugate degradation and related to the COP9-signalosome and eIF3. Cell 94, 615–623 - PubMed
-
- Leggett D. S., Glickman M. H., Finley D. (2005) Purification of proteasomes, proteasome subcomplexes, and proteasome-associated proteins from budding yeast. Methods Mol. Biol. 301, 57–70 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

