The open reading frame TTC1157 of Thermus thermophilus HB27 encodes the methyltransferase forming N²-methylguanosine at position 6 in tRNA
- PMID: 22337946
- PMCID: PMC3312568
- DOI: 10.1261/rna.030411.111
The open reading frame TTC1157 of Thermus thermophilus HB27 encodes the methyltransferase forming N²-methylguanosine at position 6 in tRNA
Abstract
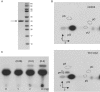
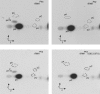
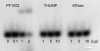
N(2)-methylguanosine (m(2)G) is found at position 6 in the acceptor stem of Thermus thermophilus tRNA(Phe). In this article, we describe the cloning, expression, and characterization of the T. thermophilus HB27 methyltransferase (MTase) encoded by the TTC1157 open reading frame that catalyzes the formation of this modified nucleoside. S-adenosyl-L-methionine is used as donor of the methyl group. The enzyme behaves as a monomer in solution. It contains an N-terminal THUMP domain predicted to bind RNA and contains a C-terminal Rossmann-fold methyltransferase (RFM) domain predicted to be responsible for catalysis. We propose to rename the TTC1157 gene trmN and the corresponding protein TrmN, according to the bacterial nomenclature of tRNA methyltransferases. Inactivation of the trmN gene in the T. thermophilus HB27 chromosome led to a total absence of m(2)G in tRNA but did not affect cell growth or the formation of other modified nucleosides in tRNA(Phe). Archaeal homologs of TrmN were identified and characterized. These proteins catalyze the same reaction as TrmN from T. thermophilus. Individual THUMP and RFM domains of PF1002 from Pyrococcus furiosus were produced. These separate domains were inactive and did not bind tRNA, reinforcing the idea that the THUMP domain acts in concert with the catalytic domain to target a particular position of the tRNA molecule.
Figures
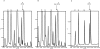



References
-
- Andersen NM, Douthwaite S 2006. YebU is a m5C methyltransferase specific for 16 S rRNA nucleotide 1407. J Mol Biol 359: 777–786 - PubMed
-
- Aravind L, Koonin EV 2001. THUMP: a predicted RNA-binding domain shared by 4-thiouridine, pseudouridine synthases and RNA methylases. Trends Biochem Sci 26: 215–217 - PubMed
-
- Armengaud J, Urbonavicius J, Fernandez B, Chaussinand G, Bujnicki JM, Grosjean H 2004. N2-methylation of guanosine at position 10 in tRNA is catalyzed by a THUMP domain-containing, S-adenosylmethionine-dependent methyltransferase, conserved in Archaea and Eukaryota. J Biol Chem 279: 37142–37152 - PubMed
-
- Auffinger P, Westhof E 1998. Location and distribution of modified nucleotides in tRNA. In Modification and editing of RNA (ed. H Grosjean, R Benne), pp. 569–576. ASM Press, Washington
-
- Awai T, Takehara T, Takeda H, Hori H 2005. Aquifex aeolicus Trm1 [tRNA (m22G26) methyltransferase] has a novel recognition mechanism of the substrate RNA. Nucleic Acids Symp Ser 49: 303–304 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
