Worm-like Ising model for protein mechanical unfolding under the effect of osmolytes
- PMID: 22339871
- PMCID: PMC3260691
- DOI: 10.1016/j.bpj.2011.12.007
Worm-like Ising model for protein mechanical unfolding under the effect of osmolytes
Abstract
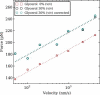
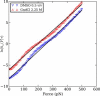
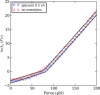
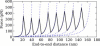
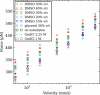
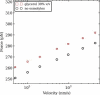
We show via single-molecule mechanical unfolding experiments that the osmolyte glycerol stabilizes the native state of the human cardiac I27 titin module against unfolding without shifting its unfolding transition state on the mechanical reaction coordinate. Taken together with similar findings on the immunoglobulin-binding domain of streptococcal protein G (GB1), these experimental results suggest that osmolytes act on proteins through a common mechanism that does not entail a shift of their unfolding transition state. We investigate the above common mechanism via an Ising-like model for protein mechanical unfolding that adds worm-like-chain behavior to a recent generalization of the Wako-Saitô-Muñoz-Eaton model with support for group-transfer free energies. The thermodynamics of the model are exactly solvable, while protein kinetics under mechanical tension can be simulated via Monte Carlo algorithms. Notably, our force-clamp and velocity-clamp simulations exhibit no shift in the position of the unfolding transition state of GB1 and I27 under the effect of various osmolytes. The excellent agreement between experiment and simulation strongly suggests that osmolytes do not assume a structural role at the mechanical unfolding transition state of proteins, acting instead by adjusting the solvent quality for the protein chain analyte.
Copyright © 2012 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
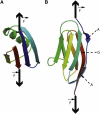

References
-
- Bolen D.W., Rose G.D. Structure and energetics of the hydrogen-bonded backbone in protein folding. Annu. Rev. Biochem. 2008;77:339–362. - PubMed
-
- Kumar S., Li M.S. Biomolecules under mechanical force. Phys. Rep. 2010;486:1–74.
-
- Aioanei D., Samorì B., Brucale M. Maximum likelihood estimation of protein kinetic parameters under weak assumptions from unfolding force spectroscopy experiments. Phys. Rev. E. 2009;80:061916. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

