Structural and genic characterization of stable genomic regions in breast cancer: relevance to chemotherapy
- PMID: 22342187
- PMCID: PMC5528331
- DOI: 10.1016/j.molonc.2012.01.001
Structural and genic characterization of stable genomic regions in breast cancer: relevance to chemotherapy
Abstract
Background: Cancer genomes accumulate frequent and diverse chromosomal abnormalities as well as gene mutations but must maintain the ability to survive in vivo. We hypothesize that genetic selection acts to maintain tumour survival by preserving copy number of specific genes and genomic regions. Genomic regions and genes that remain unaltered in copy number and expression, respectively, may be essential for maintaining tumour survival.
Methods: We analyzed copy number data of 243 previously reported breast tumours and computationally derived stable copy number regions. To identify genes in stable copy number regions with nominal changes in expression, datasets for tumour and normal samples were compared. Results were replicated by analysis of a series of independent copy number, expression and genomic sequencing studies. A subset of stable regions, including stable paralogous regions, were confirmed by quantitative PCR and fluorescence in situ hybridization (FISH) in 5 breast cancer cell lines. We deduced a comprehensive set of dually stable genes (i.e. maintaining nominal copy number and expression) which were categorized according to pathway and ontology assignments. The stability of genes encoding therapeutic drug targets was also assessed.
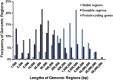
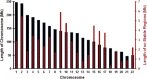
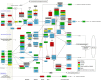
Results and conclusion: Tumour genome analysis revealed 766 unstable (amplified and/or deleted) and 812 stable contiguous genomic regions. Replication analysis of an independent set of 171 breast tumours confirmed copy number stability of 1.3 Gb of the genome. We found that 5804 of these genes were dually stable. The composition of this gene set remained essentially unchanged (<2% reduction) after accounting for commonly mutated breast cancer genes found by sequencing and differential expression. The stable breast cancer genome is enriched for cellular metabolism, regulation of gene expression, DNA packaging (chromatin and nucleosome assembly), and regulation of apoptosis functions. Stable genes participating in multiple essential pathways were consistently found to be targets of chemotherapies. Preservation of stable, essential genes may be related to the effectiveness of certain chemotherapeutic agents that act on multiple gene products in this set.
Copyright © 2012 Federation of European Biochemical Societies. Published by Elsevier B.V. All rights reserved.
Figures

Similar articles
-
Comprehensive copy number profiles of breast cancer cell model genomes.Breast Cancer Res. 2006;8(1):R9. doi: 10.1186/bcr1370. Epub 2006 Jan 3. Breast Cancer Res. 2006. PMID: 16417655 Free PMC article.
-
Copy number analysis of c-erb-B2 (HER-2/neu) and topoisomerase IIalpha genes in breast carcinoma by quantitative real-time polymerase chain reaction using hybridization probes and fluorescence in situ hybridization.Arch Pathol Lab Med. 2005 Jan;129(1):39-46. doi: 10.5858/2005-129-39-CNAOCN. Arch Pathol Lab Med. 2005. PMID: 15628907
-
Genomic profiling by array comparative genomic hybridization reveals novel DNA copy number changes in breast phyllodes tumours.Cell Oncol. 2009;31(1):31-9. doi: 10.3233/clo-2009-0457. Cell Oncol. 2009. PMID: 19096148 Free PMC article.
-
Genome-wide copy number analysis in primary breast cancer.Expert Opin Ther Targets. 2012 Mar;16 Suppl 1:S31-5. doi: 10.1517/14728222.2011.636739. Epub 2012 Feb 8. Expert Opin Ther Targets. 2012. PMID: 22313367 Review.
-
The genomic landscape of breast cancer brain metastases: a systematic review.Lancet Oncol. 2021 Jan;22(1):e7-e17. doi: 10.1016/S1470-2045(20)30556-8. Lancet Oncol. 2021. PMID: 33387511
Cited by
-
A High-Quality Blue Whale Genome, Segmental Duplications, and Historical Demography.Mol Biol Evol. 2024 Mar 1;41(3):msae036. doi: 10.1093/molbev/msae036. Mol Biol Evol. 2024. PMID: 38376487 Free PMC article.
-
Recent Advances in Discovering the Role of CCL5 in Metastatic Breast Cancer.Mini Rev Med Chem. 2015;15(13):1063-72. doi: 10.2174/138955751513150923094709. Mini Rev Med Chem. 2015. PMID: 26420723 Free PMC article. Review.
-
Relationship between drug targets and drug-signature networks: a network-based genome-wide landscape.BMC Med Genomics. 2023 Jan 30;16(1):17. doi: 10.1186/s12920-023-01444-8. BMC Med Genomics. 2023. PMID: 36717817 Free PMC article.
-
Next Generation Sequencing of Circulating Cell-Free DNA for Evaluating Mutations and Gene Amplification in Metastatic Breast Cancer.Clin Chem. 2017 Feb;63(2):532-541. doi: 10.1373/clinchem.2016.261834. Epub 2016 Dec 9. Clin Chem. 2017. PMID: 27940449 Free PMC article.
-
Construction of competing endogenous RNA networks from paired RNA-seq data sets by pointwise mutual information.BMC Genomics. 2019 Dec 24;20(Suppl 9):943. doi: 10.1186/s12864-019-6321-x. BMC Genomics. 2019. PMID: 31874629 Free PMC article.
References
-
- Ataee, R. , Ajdary, S. , Zarrindast, M. , Rezayat, M. , Shokrgozar, M.A. , Ataee, A. , 2010. Y25130 hydrochloride, a selective 5HT3 receptor antagonist has potent antimitogenic and apoptotic effect on HT29 colorectal cancer cell line. Eur. J. Cancer Prev.. 19, 138–143. - PubMed
-
- Chin, S.F. , Teschendorff, A.E. , Marioni, J.C. , Wang, Y. , Barbosa-Morais, N.L. , Thorne, N.P. , Costa, J.L. , Pinder, S.E. , van de Wiel, M.A. , Green, A.R. , 2007. High-resolution aCGH and expression profiling identifies a novel genomic subtype of ER negative breast cancer. Genome Biol.. 8, R215 - PMC - PubMed
-
- Clynes, R.A. , Towers, T.L. , Presta, L.G. , Ravetch, J.V. , 2000. Inhibitory Fc receptors modulate in vivo cytoxicity against tumor targets. Nat. Med.. 6, 443–446. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

