The circadian clock controls toll-like receptor 9-mediated innate and adaptive immunity
- PMID: 22342842
- PMCID: PMC3315694
- DOI: 10.1016/j.immuni.2011.12.017
The circadian clock controls toll-like receptor 9-mediated innate and adaptive immunity
Abstract
Circadian rhythms refer to biologic processes that oscillate with a period of ~24 hr. These rhythms are sustained by a molecular clock and provide a temporal matrix that ensures the coordination of homeostatic processes with the periodicity of environmental challenges. We demonstrate the circadian molecular clock controls the expression and function of Toll-like receptor 9 (TLR9). In a vaccination model using TLR9 ligand as adjuvant, mice immunized at the time of enhanced TLR9 responsiveness presented weeks later with an improved adaptive immune response. In a TLR9-dependent mouse model of sepsis, we found that disease severity was dependent on the timing of sepsis induction, coinciding with the daily changes in TLR9 expression and function. These findings unveil a direct molecular link between the circadian and innate immune systems with important implications for immunoprophylaxis and immunotherapy.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures

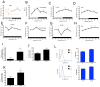
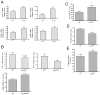

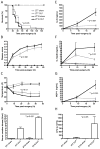
Comment in
-
Toll-like receptor 9, what o'clock is it?Immunity. 2012 Feb 24;36(2):159-61. doi: 10.1016/j.immuni.2012.02.003. Immunity. 2012. PMID: 22365660
-
Gene regulation: Clocking on for TLR9.Nat Rev Immunol. 2012 Mar 9;12(4):228-9. doi: 10.1038/nri3194. Nat Rev Immunol. 2012. PMID: 22402671 No abstract available.
-
Leukocyte: tempus fugit vel carpe diem.Am J Transplant. 2012 Jul;12(7):1665. doi: 10.1111/j.1600-6143.2012.04187.x. Am J Transplant. 2012. PMID: 22741751 No abstract available.
Similar articles
-
Gene regulation: Clocking on for TLR9.Nat Rev Immunol. 2012 Mar 9;12(4):228-9. doi: 10.1038/nri3194. Nat Rev Immunol. 2012. PMID: 22402671 No abstract available.
-
Circadian clock components Bmal1 and Clock1 regulate tlr9 gene expression in the Japanese medaka (Oryzias latipes).Fish Shellfish Immunol. 2020 Oct;105:438-445. doi: 10.1016/j.fsi.2020.07.009. Epub 2020 Jul 9. Fish Shellfish Immunol. 2020. PMID: 32653586
-
Daily variation in macrophage phagocytosis is clock-independent and dispensable for cytokine production.Immunology. 2019 Jun;157(2):122-136. doi: 10.1111/imm.13053. Epub 2019 Apr 16. Immunology. 2019. PMID: 30773630 Free PMC article.
-
Clocking in to immunity.Nat Rev Immunol. 2018 Jul;18(7):423-437. doi: 10.1038/s41577-018-0008-4. Nat Rev Immunol. 2018. PMID: 29662121 Review.
-
Circadian Clocks in the Immune System.J Biol Rhythms. 2015 Aug;30(4):277-90. doi: 10.1177/0748730415577723. Epub 2015 Apr 20. J Biol Rhythms. 2015. PMID: 25900041 Review.
Cited by
-
Circadian Effects of Drug Responses.Annu Rev Biomed Eng. 2021 Jul 13;23:203-224. doi: 10.1146/annurev-bioeng-082120-034725. Epub 2021 Mar 31. Annu Rev Biomed Eng. 2021. PMID: 33788580 Free PMC article.
-
A ticking clock links metabolic pathways and organ systems function in health and disease.Clin Exp Med. 2014 May;14(2):133-40. doi: 10.1007/s10238-013-0235-8. Epub 2013 Apr 4. Clin Exp Med. 2014. PMID: 23553127 Review.
-
The Metronome of Symbiosis: Interactions Between Microbes and the Host Circadian Clock.Integr Comp Biol. 2016 Nov;56(5):776-783. doi: 10.1093/icb/icw067. Epub 2016 Jul 1. Integr Comp Biol. 2016. PMID: 27371387 Free PMC article.
-
Do macrophages follow the beat of circadian rhythm in TIME (Tumor Immune Microenvironment)?F1000Res. 2023 Jan 27;12:101. doi: 10.12688/f1000research.129863.1. eCollection 2023. F1000Res. 2023. PMID: 37469718 Free PMC article. Review.
-
Bacterial surface colonization, preferential attachment and fitness under periodic stress.PLoS Comput Biol. 2019 Mar 5;15(3):e1006815. doi: 10.1371/journal.pcbi.1006815. eCollection 2019 Mar. PLoS Comput Biol. 2019. PMID: 30835727 Free PMC article.
References
-
- Akira S, Takeda K, Kaisho T. Toll-like receptors: critical proteins linking innate and acquired immunity. Nature immunology. 2001;2:675–680. - PubMed
-
- Arjona A, Sarkar DK. Circadian oscillations of clock genes, cytolytic factors, and cytokines in rat NK cells. J Immunol. 2005;174:7618–7624. - PubMed
-
- Arjona A, Sarkar DK. The circadian gene mPer2 regulates the daily rhythm of IFN-gamma. J Interferon Cytokine Res. 2006;26:645–649. - PubMed
-
- Balsalobre A, Damiola F, Schibler U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell. 1998;93:929–937. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

