Separate regions of glucocorticoid receptor, coactivator TIF2, and comodulator STAMP modify different parameters of glucocorticoid-mediated gene induction
- PMID: 22342989
- PMCID: PMC3312974
- DOI: 10.1016/j.mce.2012.02.001
Separate regions of glucocorticoid receptor, coactivator TIF2, and comodulator STAMP modify different parameters of glucocorticoid-mediated gene induction
Abstract
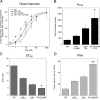
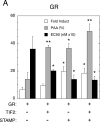
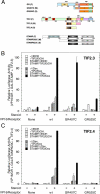
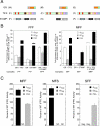
Increased specificity in steroid-regulated gene expression is a long-sought goal of endocrinologists. Considerable progress has resulted from the discovery of coactivators, corepressors, and comodulators that adjust the total activity (A(max)) of gene induction. Two less frequently quantitated, but equally potent, means of improving specificity are the concentration of agonist steroid required for half-maximal activity (EC(50)) and the residual or partial agonist activity displayed by most antisteroids (PAA). It is usually assumed that the modulatory activity of transcriptional cofactors coordinately regulates A(max), EC(50), and PAA. Here we examine the hypothesis that these three parameters can be independently modified by separate protein domains. The test system involves three differently sized fragments of each of three factors (glucocorticoid receptor [GR], coactivator TIF2, and comodulator STAMP), which are shown to form a ternary complex and similarly affect the induction properties of transfected and endogenous genes. Twenty-five different fragment combinations of the ternary complex are examined for their ability to modulate the A(max), EC(50), and PAA of a transiently transfected synthetic reporter gene. Different combinations selectively alter one, two, or all three parameters. These results clearly demonstrate that A(max), EC(50), and PAA can be independently regulated under some conditions by different pathways or molecular interactions. This new mechanistic insight suggests that selected activities of individual transcription factors are attractive targets for small molecules, which would have obvious clinical applications for increasing the specificity of steroids during endocrine therapies.
Published by Elsevier Ireland Ltd.
Figures





Similar articles
-
Equilibrium interactions of corepressors and coactivators with agonist and antagonist complexes of glucocorticoid receptors.Mol Endocrinol. 2004 Jun;18(6):1376-95. doi: 10.1210/me.2003-0421. Epub 2004 Mar 11. Mol Endocrinol. 2004. PMID: 15016838
-
Opposing effects of corepressor and coactivators in determining the dose-response curve of agonists, and residual agonist activity of antagonists, for glucocorticoid receptor-regulated gene expression.Mol Endocrinol. 1999 Dec;13(12):2108-21. doi: 10.1210/mend.13.12.0384. Mol Endocrinol. 1999. PMID: 10598585
-
Modulation of transcription parameters in glucocorticoid receptor-mediated repression.Mol Cell Endocrinol. 2008 Nov 25;295(1-2):59-69. doi: 10.1016/j.mce.2008.05.008. Epub 2008 May 21. Mol Cell Endocrinol. 2008. PMID: 18583028 Free PMC article.
-
Glucocorticoid receptor cofactors as therapeutic targets.Curr Opin Pharmacol. 2010 Dec;10(6):613-9. doi: 10.1016/j.coph.2010.08.001. Epub 2010 Aug 26. Curr Opin Pharmacol. 2010. PMID: 20801081 Free PMC article. Review.
-
The road less traveled: new views of steroid receptor action from the path of dose-response curves.Mol Cell Endocrinol. 2012 Jan 30;348(2):373-82. doi: 10.1016/j.mce.2011.05.030. Epub 2011 Jun 1. Mol Cell Endocrinol. 2012. PMID: 21664235 Free PMC article. Review.
Cited by
-
Disruption of Ttll5/stamp gene (tubulin tyrosine ligase-like protein 5/SRC-1 and TIF2-associated modulatory protein gene) in male mice causes sperm malformation and infertility.J Biol Chem. 2013 May 24;288(21):15167-80. doi: 10.1074/jbc.M113.453936. Epub 2013 Apr 4. J Biol Chem. 2013. PMID: 23558686 Free PMC article.
-
Kinetically-defined component actions in gene repression.PLoS Comput Biol. 2015 Mar 27;11(3):e1004122. doi: 10.1371/journal.pcbi.1004122. eCollection 2015 Mar. PLoS Comput Biol. 2015. PMID: 25816223 Free PMC article.
-
PA1 protein, a new competitive decelerator acting at more than one step to impede glucocorticoid receptor-mediated transactivation.J Biol Chem. 2013 Jan 4;288(1):42-58. doi: 10.1074/jbc.M112.427740. Epub 2012 Nov 17. J Biol Chem. 2013. PMID: 23161582 Free PMC article.
-
Minireview: dynamic structures of nuclear hormone receptors: new promises and challenges.Mol Endocrinol. 2014 Feb;28(2):173-82. doi: 10.1210/me.2013-1334. Epub 2013 Nov 27. Mol Endocrinol. 2014. PMID: 24284822 Free PMC article. Review.
-
Research resource: modulators of glucocorticoid receptor activity identified by a new high-throughput screening assay.Mol Endocrinol. 2014 Jul;28(7):1194-206. doi: 10.1210/me.2014-1069. Epub 2014 May 21. Mol Endocrinol. 2014. PMID: 24850414 Free PMC article.
References
-
- Avenant C, Kotitschke A, Hapgood JP. Glucocorticoid receptor phosphorylation modulates transcription efficacy through GRIP-1 recruitment. Biochemistry. 2010;49:972–985. - PubMed
-
- Chen W, Rogatsky I, Garabedian MJ. MED14 and MED1 differentially regulate target-specific gene activation by the glucocorticoid receptor. Mol Endocrinol. 2006;20:560–572. - PubMed
-
- Cho S, Blackford JA, Jr., Simons SS., Jr. Role of activation function domain 1, DNA binding, and coactivator in the expression of partial agonist activity of glucocorticoid receptor complexes. Biochemistry. 2005a;44:3547–3561. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

