Geobacillus thermodenitrificans YjbH recognizes the C-terminal end of Bacillus subtilis Spx to accelerate Spx proteolysis by ClpXP
- PMID: 22343351
- PMCID: PMC3542823
- DOI: 10.1099/mic.0.057661-0
Geobacillus thermodenitrificans YjbH recognizes the C-terminal end of Bacillus subtilis Spx to accelerate Spx proteolysis by ClpXP
Abstract
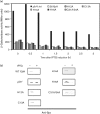
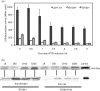
Proteolytic control can govern the levels of specific regulatory factors, such as Spx, a transcriptional regulator of the oxidative stress response in Gram-positive bacteria. Under oxidative stress, Spx concentration is elevated and upregulates transcription of genes that function in the stress response. When stress is alleviated, proteolysis of Spx catalysed by ClpXP reduces Spx concentration. Proteolysis is enhanced by the substrate recognition factor YjbH, which possesses a His-Cys-rich region at its N terminus. However, mutations that generate H12A, C13A, H14A, H16A and C31/34A residue substitutions in the N terminus of Bacillus subtilis YjbH (BsYjbH) do not affect functionality in Spx proteolytic control in vivo and in vitro. Because of difficulties in obtaining soluble BsYjbH, the Geobacillus thermodenitrificans yjbH gene was cloned, which yielded soluble GtYjbH protein. Despite its lack of a His-Cys-rich region, GtYjbH complements a B. subtilis yjbH null mutant, and shows high activity in vitro when combined with ClpXP and Spx in an approximately 30 : 1 (ClpXP/Spx : GtYjbH) molar ratio. In vitro interaction experiments showed that Spx and the protease-resistant Spx(DD) (in which the last two residues of Spx are replaced with two Asp residues) bind to GtYjbH, but deletion of 12 residues from the Spx C terminus (SpxΔC) significantly diminished interaction and proteolytic degradation, indicating that the C terminus of Spx is important for YjbH recognition. These experiments also showed that Spx, but not GtYjbH, interacts with ClpX. Kinetic measurements for Spx proteolysis by ClpXP in the presence and absence of GtYjbH suggest that YjbH overcomes non-productive Spx-ClpX interaction, resulting in rapid degradation.
Figures





Similar articles
-
Adaptor bypass mutations of Bacillus subtilis spx suggest a mechanism for YjbH-enhanced proteolysis of the regulator Spx by ClpXP.Mol Microbiol. 2014 Aug;93(3):426-38. doi: 10.1111/mmi.12671. Epub 2014 Jul 10. Mol Microbiol. 2014. PMID: 24942655 Free PMC article.
-
The YjbH protein of Bacillus subtilis enhances ClpXP-catalyzed proteolysis of Spx.J Bacteriol. 2009 Feb;191(4):1268-77. doi: 10.1128/JB.01289-08. Epub 2008 Dec 12. J Bacteriol. 2009. PMID: 19074380 Free PMC article.
-
The YjbH adaptor protein enhances proteolysis of the transcriptional regulator Spx in Staphylococcus aureus.J Bacteriol. 2012 Mar;194(5):1186-94. doi: 10.1128/JB.06414-11. Epub 2011 Dec 22. J Bacteriol. 2012. PMID: 22194450 Free PMC article.
-
Roles and regulation of Spx family transcription factors in Bacillus subtilis and related species.Adv Microb Physiol. 2019;75:279-323. doi: 10.1016/bs.ampbs.2019.05.003. Epub 2019 Jul 5. Adv Microb Physiol. 2019. PMID: 31655740 Free PMC article. Review.
-
Spx, a versatile regulator of the Bacillus subtilis stress response.Curr Genet. 2019 Aug;65(4):871-876. doi: 10.1007/s00294-019-00950-6. Epub 2019 Mar 4. Curr Genet. 2019. PMID: 30830258 Review.
Cited by
-
YjbH Solubility Controls Spx in Staphylococcus aureus: Implication for MazEF Toxin-Antitoxin System Regulation.Front Microbiol. 2020 Feb 6;11:113. doi: 10.3389/fmicb.2020.00113. eCollection 2020. Front Microbiol. 2020. PMID: 32117138 Free PMC article.
-
YjbH Requires Its Thioredoxin Active Motif for the Nitrosative Stress Response, Cell-to-Cell Spread, and Protein-Protein Interactions in Listeria monocytogenes.J Bacteriol. 2020 May 27;202(12):e00099-20. doi: 10.1128/JB.00099-20. Print 2020 May 27. J Bacteriol. 2020. PMID: 32253340 Free PMC article.
-
Role of adaptor TrfA and ClpPC in controlling levels of SsrA-tagged proteins and antitoxins in Staphylococcus aureus.J Bacteriol. 2014 Dec;196(23):4140-51. doi: 10.1128/JB.02222-14. Epub 2014 Sep 15. J Bacteriol. 2014. PMID: 25225270 Free PMC article.
-
Update on the Protein Homeostasis Network in Bacillus subtilis.Front Microbiol. 2022 Mar 8;13:865141. doi: 10.3389/fmicb.2022.865141. eCollection 2022. Front Microbiol. 2022. PMID: 35350626 Free PMC article. Review.
-
Thiol-based redox switches in prokaryotes.Biol Chem. 2015 May;396(5):415-44. doi: 10.1515/hsz-2015-0102. Biol Chem. 2015. PMID: 25720121 Free PMC article. Review.
References
-
- Charbonnier Y., Gettler B., François P., Bento M., Renzoni A., Vaudaux P., Schlegel W., Schrenzel J. (2005). A generic approach for the design of whole-genome oligoarrays, validated for genomotyping, deletion mapping and gene expression analysis on Staphylococcus aureus. BMC Genomics 6, 95. 10.1186/1471-2164-6-95 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

