The TPR domain in the host Cyp40-like cyclophilin binds to the viral replication protein and inhibits the assembly of the tombusviral replicase
- PMID: 22346747
- PMCID: PMC3276564
- DOI: 10.1371/journal.ppat.1002491
The TPR domain in the host Cyp40-like cyclophilin binds to the viral replication protein and inhibits the assembly of the tombusviral replicase
Abstract
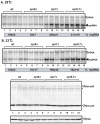
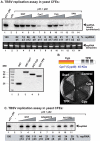
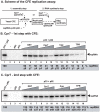
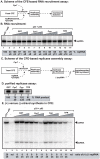
Replication of plus-stranded RNA viruses is greatly affected by numerous host-coded proteins acting either as susceptibility or resistance factors. Previous genome-wide screens and global proteomics approaches with Tomato bushy stunt tombusvirus (TBSV) in a yeast model host revealed the involvement of cyclophilins, which are a large family of host prolyl isomerases, in TBSV replication. In this paper, we identified those members of the large cyclophilin family that interacted with the viral replication proteins and inhibited TBSV replication. Further characterization of the most effective cyclophilin, the Cyp40-like Cpr7p, revealed that it strongly inhibits many steps during TBSV replication in a cell-free replication assay. These steps include viral RNA recruitment inhibited via binding of Cpr7p to the RNA-binding region of the viral replication protein; the assembly of the viral replicase complex and viral RNA synthesis. Since the TPR (tetratricopeptide repeats) domain, but not the catalytic domain of Cpr7p is needed for the inhibitory effect on TBSV replication, it seems that the chaperone activity of Cpr7p provides the negative regulatory function. We also show that three Cyp40-like proteins from plants can inhibit TBSV replication in vitro and Cpr7p is also effective against Nodamura virus, an insect pathogen. Overall, the current work revealed a role for Cyp40-like proteins and their TPR domains as regulators of RNA virus replication.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures





Similar articles
-
Cyclophilin A binds to the viral RNA and replication proteins, resulting in inhibition of tombusviral replicase assembly.J Virol. 2013 Dec;87(24):13330-42. doi: 10.1128/JVI.02101-13. Epub 2013 Oct 2. J Virol. 2013. PMID: 24089553 Free PMC article.
-
Identification of novel host factors via conserved domain search: Cns1 cochaperone is a novel restriction factor of tombusvirus replication in yeast.J Virol. 2013 Dec;87(23):12600-10. doi: 10.1128/JVI.00196-13. Epub 2013 Sep 11. J Virol. 2013. PMID: 24027337 Free PMC article.
-
Cpr1 cyclophilin and Ess1 parvulin prolyl isomerases interact with the tombusvirus replication protein and inhibit viral replication in yeast model host.Virology. 2010 Oct 25;406(2):342-51. doi: 10.1016/j.virol.2010.07.022. Epub 2010 Aug 14. Virology. 2010. PMID: 20709345
-
Global genomics and proteomics approaches to identify host factors as targets to induce resistance against Tomato bushy stunt virus.Adv Virus Res. 2010;76:123-77. doi: 10.1016/S0065-3527(10)76004-8. Epub 2010 Mar 31. Adv Virus Res. 2010. PMID: 20965073 Free PMC article. Review.
-
The roles of host factors in tombusvirus RNA recombination.Adv Virus Res. 2011;81:63-84. doi: 10.1016/B978-0-12-385885-6.00008-0. Adv Virus Res. 2011. PMID: 22094079 Review.
Cited by
-
Authentic in vitro replication of two tombusviruses in isolated mitochondrial and endoplasmic reticulum membranes.J Virol. 2012 Dec;86(23):12779-94. doi: 10.1128/JVI.00973-12. Epub 2012 Sep 12. J Virol. 2012. PMID: 22973028 Free PMC article.
-
Race against Time between the Virus and Host: Actin-Assisted Rapid Biogenesis of Replication Organelles is Used by TBSV to Limit the Recruitment of Cellular Restriction Factors.J Virol. 2022 Jun 22;96(12):e0016821. doi: 10.1128/jvi.00168-21. Epub 2022 May 31. J Virol. 2022. PMID: 35638821 Free PMC article.
-
Cyclophilin A binds to the viral RNA and replication proteins, resulting in inhibition of tombusviral replicase assembly.J Virol. 2013 Dec;87(24):13330-42. doi: 10.1128/JVI.02101-13. Epub 2013 Oct 2. J Virol. 2013. PMID: 24089553 Free PMC article.
-
Proteome-wide overexpression of host proteins for identification of factors affecting tombusvirus RNA replication: an inhibitory role of protein kinase C.J Virol. 2012 Sep;86(17):9384-95. doi: 10.1128/JVI.00019-12. Epub 2012 Jun 20. J Virol. 2012. PMID: 22718827 Free PMC article.
-
Identification of giant Mimivirus protein functions using RNA interference.Front Microbiol. 2015 Apr 28;6:345. doi: 10.3389/fmicb.2015.00345. eCollection 2015. Front Microbiol. 2015. PMID: 25972846 Free PMC article.
References
-
- Nagy PD. Yeast as a model host to explore plant virus-host interactions. Annu Rev Phytopathol. 2008;46:217–242. - PubMed
-
- Nagy PD, Pogany J. Multiple roles of viral replication proteins in plant RNA virus replication. Methods Mol Biol. 2008;451:55–68. - PubMed
-
- Nagy PD, Wang RY, Pogany J, Hafren A, Makinen K. Emerging picture of host chaperone and cyclophilin roles in RNA virus replication. Virology. 2011;411:374–382. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

