Infection Strategies of Bacterial and Viral Pathogens through Pathogen-Human Protein-Protein Interactions
- PMID: 22347880
- PMCID: PMC3278985
- DOI: 10.3389/fmicb.2012.00046
Infection Strategies of Bacterial and Viral Pathogens through Pathogen-Human Protein-Protein Interactions
Abstract
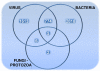
Since ancient times, even in today's modern world, infectious diseases cause lots of people to die. Infectious organisms, pathogens, cause diseases by physical interactions with human proteins. A thorough analysis of these interspecies interactions is required to provide insights about infection strategies of pathogens. Here we analyzed the most comprehensive available pathogen-human protein interaction data including 23,435 interactions, targeting 5,210 human proteins. The data were obtained from the newly developed pathogen-host interaction search tool, PHISTO. This is the first comprehensive attempt to get a comparison between bacterial and viral infections. We investigated human proteins that are targeted by bacteria and viruses to provide an overview of common and special infection strategies used by these pathogen types. We observed that in the human protein interaction network the proteins targeted by pathogens have higher connectivity and betweenness centrality values than those proteins not interacting with pathogens. The preference of interacting with hub and bottleneck proteins is found to be a common infection strategy of all types of pathogens to manipulate essential mechanisms in human. Compared to bacteria, viruses tend to interact with human proteins of much higher connectivity and centrality values in the human network. Gene Ontology enrichment analysis of the human proteins targeted by pathogens indicates crucial clues about the infection mechanisms of bacteria and viruses. As the main infection strategy, bacteria interact with human proteins that function in immune response to disrupt human defense mechanisms. Indispensable viral strategy, on the other hand, is the manipulation of human cellular processes in order to use that transcriptional machinery for their own genetic material transcription. A novel observation about pathogen-human systems is that the human proteins targeted by both pathogens are enriched in the regulation of metabolic processes.
Keywords: PHISTO; bottleneck; gene ontology; hub; infection strategy; pathogen–human protein–protein interactions.
Figures
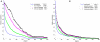


Similar articles
-
The landscape of human proteins interacting with viruses and other pathogens.PLoS Pathog. 2008 Feb 8;4(2):e32. doi: 10.1371/journal.ppat.0040032. PLoS Pathog. 2008. PMID: 18282095 Free PMC article.
-
Comparative interactomics for virus-human protein-protein interactions: DNA viruses versus RNA viruses.FEBS Open Bio. 2017 Jan 4;7(1):96-107. doi: 10.1002/2211-5463.12167. eCollection 2017 Jan. FEBS Open Bio. 2017. PMID: 28097092 Free PMC article.
-
PHISTO: pathogen-host interaction search tool.Bioinformatics. 2013 May 15;29(10):1357-8. doi: 10.1093/bioinformatics/btt137. Epub 2013 Mar 20. Bioinformatics. 2013. PMID: 23515528
-
Analysis of protein targets in pathogen-host interaction in infectious diseases: a case study on Plasmodium falciparum and Homo sapiens interaction network.Brief Funct Genomics. 2018 Nov 26;17(6):441-450. doi: 10.1093/bfgp/elx024. Brief Funct Genomics. 2018. PMID: 29028886 Review.
-
How Viral and Intracellular Bacterial Pathogens Reprogram the Metabolism of Host Cells to Allow Their Intracellular Replication.Front Cell Infect Microbiol. 2019 Mar 4;9:42. doi: 10.3389/fcimb.2019.00042. eCollection 2019. Front Cell Infect Microbiol. 2019. PMID: 30886834 Free PMC article. Review.
Cited by
-
Mining host-pathogen protein interactions to characterize Burkholderia mallei infectivity mechanisms.PLoS Comput Biol. 2015 Mar 4;11(3):e1004088. doi: 10.1371/journal.pcbi.1004088. eCollection 2015 Mar. PLoS Comput Biol. 2015. PMID: 25738731 Free PMC article.
-
The domain landscape of virus-host interactomes.Biomed Res Int. 2014;2014:867235. doi: 10.1155/2014/867235. Epub 2014 Jun 4. Biomed Res Int. 2014. PMID: 24991570 Free PMC article.
-
Science-Based Strategies of Antiviral Coatings with Viricidal Properties for the COVID-19 Like Pandemics.Materials (Basel). 2020 Sep 11;13(18):4041. doi: 10.3390/ma13184041. Materials (Basel). 2020. PMID: 32933043 Free PMC article. Review.
-
Annexin A2 in Inflammation and Host Defense.Cells. 2020 Jun 19;9(6):1499. doi: 10.3390/cells9061499. Cells. 2020. PMID: 32575495 Free PMC article. Review.
-
Techniques for transferring host-pathogen protein interactions knowledge to new tasks.Front Microbiol. 2015 Feb 2;6:36. doi: 10.3389/fmicb.2015.00036. eCollection 2015. Front Microbiol. 2015. PMID: 25699028 Free PMC article.
References
-
- Ashburner M., Ball C. A., Blake J. A., Botstein D., Butler H., Cherry M., Davis A. P., Dolinski K., Dwight S. S., Eppig J. T., Harris M. A., Hill D. P., Issel-Tarver L., Kasarskis A., Lewis S., Matese J. C., Richardson J. E., Ringwald M., Rubin G. M., Sherlock G. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–2910.1038/75556 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Research Materials

