Multilocus microsatellite typing (MLMT) of strains from Turkey and Cyprus reveals a novel monophyletic L. donovani sensu lato group
- PMID: 22348162
- PMCID: PMC3279343
- DOI: 10.1371/journal.pntd.0001507
Multilocus microsatellite typing (MLMT) of strains from Turkey and Cyprus reveals a novel monophyletic L. donovani sensu lato group
Abstract
Background: New foci of human CL caused by strains of the Leishmania donovani (L. donovani) complex have been recently described in Cyprus and the Çukurova region in Turkey (L. infantum) situated 150 km north of Cyprus. Cypriot strains were typed by Multilocus Enzyme Electrophoresis (MLEE) using the Montpellier (MON) system as L. donovani zymodeme MON-37. However, multilocus microsatellite typing (MLMT) has shown that this zymodeme is paraphyletic; composed of distantly related genetic subgroups of different geographical origin. Consequently the origin of the Cypriot strains remained enigmatic.
Methodology/principal findings: The Cypriot strains were compared with a set of Turkish isolates obtained from a CL patient and sand fly vectors in south-east Turkey (Çukurova region; CUK strains) and from a VL patient in the south-west (Kuşadasi; EP59 strain). These Turkish strains were initially analyzed using the K26-PCR assay that discriminates MON-1 strains by their amplicon size. In line with previous DNA-based data, the strains were inferred to the L. donovani complex and characterized as non MON-1. For these strains MLEE typing revealed two novel zymodemes; L. donovani MON-309 (CUK strains) and MON-308 (EP59). A population genetic analysis of the Turkish isolates was performed using 14 hyper-variable microsatellite loci. The genotypic profiles of 68 previously analyzed L. donovani complex strains from major endemic regions were included for comparison. Population structures were inferred by combination of bayesian model-based and distance-based approaches. MLMT placed the Turkish and Cypriot strains in a subclade of a newly discovered, genetically distinct L. infantum monophyletic group, suggesting that the Cypriot strains may originate from Turkey.
Conclusion: The discovery of a genetically distinct L. infantum monophyletic group in the south-eastern Mediterranean stresses the importance of species genetic characterization towards better understanding, monitoring and controlling the spread of leishmaniasis in this region.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

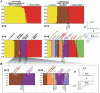


References
-
- WHO. 2010. Control of leishmaniasis: report of a meeting of the WHO expert committee on the control of Leishmaniases, Geneva, 22–26 March 2010.
-
- Gramiccia M, Gradoni L, Troiani M. Heterogeneity among zymodemes of Leishmania infantum from HIV-positive patients with visceral leishmaniasis in south Italy. FEMS microbiology letters. 1995;128:33–38. - PubMed
-
- Pratlong F, Dedet JP, Marty P, Portus M, Deniau M, et al. Leishmania-human immunodeficiency virus coinfection in the Mediterranean basin: isoenzymatic characterization of 100 isolates of the Leishmania infantum complex. J Infect Dis. 1995;172:323–326. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

