Endogenous voltage gradients as mediators of cell-cell communication: strategies for investigating bioelectrical signals during pattern formation
- PMID: 22350846
- PMCID: PMC3869965
- DOI: 10.1007/s00441-012-1329-4
Endogenous voltage gradients as mediators of cell-cell communication: strategies for investigating bioelectrical signals during pattern formation
Abstract
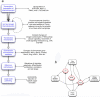
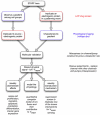
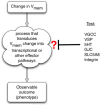
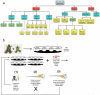
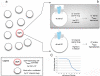
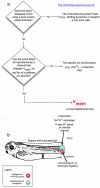
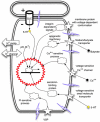
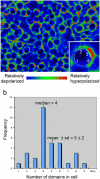
Alongside the well-known chemical modes of cell-cell communication, we find an important and powerful system of bioelectrical signaling: changes in the resting voltage potential (Vmem) of the plasma membrane driven by ion channels, pumps and gap junctions. Slow Vmem changes in all cells serve as a highly conserved, information-bearing pathway that regulates cell proliferation, migration and differentiation. In embryonic and regenerative pattern formation and in the disorganization of neoplasia, bioelectrical cues serve as mediators of large-scale anatomical polarity, organ identity and positional information. Recent developments have resulted in tools that enable a high-resolution analysis of these biophysical signals and their linkage with upstream and downstream canonical genetic pathways. Here, we provide an overview for the study of bioelectric signaling, focusing on state-of-the-art approaches that use molecular physiology and developmental genetics to probe the roles of bioelectric events functionally. We highlight the logic, strategies and well-developed technologies that any group of researchers can employ to identify and dissect ionic signaling components in their own work and thus to help crack the bioelectric code. The dissection of bioelectric events as instructive signals enabling the orchestration of cell behaviors into large-scale coherent patterning programs will enrich on-going work in diverse areas of biology, as biophysical factors become incorporated into our systems-level understanding of cell interactions.
Figures


References
-
- Adams DS. A new tool for tissue engineers: ions as regulators of morphogenesis during development and regeneration. Tissue Eng. 2008;14:1461–1468. - PubMed
-
- Adams DS, Levin M. Strategies and techniques for investigation of biophysical signals in patterning. In: Whitman M, Sater AK, editors. Analysis of growth factor signaling in embryos. CRC Press; Boca Raton: 2006b. pp. 177–262.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

