Transcriptome characterization and sequencing-based identification of salt-responsive genes in Millettia pinnata, a semi-mangrove plant
- PMID: 22351699
- PMCID: PMC3325082
- DOI: 10.1093/dnares/dss004
Transcriptome characterization and sequencing-based identification of salt-responsive genes in Millettia pinnata, a semi-mangrove plant
Abstract
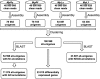
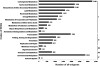
Semi-mangroves form a group of transitional species between glycophytes and halophytes, and hold unique potential for learning molecular mechanisms underlying plant salt tolerance. Millettia pinnata is a semi-mangrove plant that can survive a wide range of saline conditions in the absence of specialized morphological and physiological traits. By employing the Illumina sequencing platform, we generated ~192 million short reads from four cDNA libraries of M. pinnata and processed them into 108,598 unisequences with a high depth of coverage. The mean length and total length of these unisequences were 606 bp and 65.8 Mb, respectively. A total of 54,596 (50.3%) unisequences were assigned Nr annotations. Functional classification revealed the involvement of unisequences in various biological processes related to metabolism and environmental adaptation. We identified 23,815 candidate salt-responsive genes with significantly differential expression under seawater and freshwater treatments. Based on the reverse transcription-polymerase chain reaction (RT-PCR) and real-time PCR analyses, we verified the changes in expression levels for a number of candidate genes. The functional enrichment analyses for the candidate genes showed tissue-specific patterns of transcriptome remodelling upon salt stress in the roots and the leaves. The transcriptome of M. pinnata will provide valuable gene resources for future application in crop improvement. In addition, this study sets a good example for large-scale identification of salt-responsive genes in non-model organisms using the sequencing-based approach.
Figures
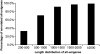
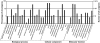

Similar articles
-
Isolation and Functional Characterization of a Salt-Responsive Calmodulin-Like Gene MpCML40 from Semi-Mangrove Millettia pinnata.Int J Mol Sci. 2021 Mar 27;22(7):3475. doi: 10.3390/ijms22073475. Int J Mol Sci. 2021. PMID: 33801703 Free PMC article.
-
Transcriptome analysis of grapevine under salinity and identification of key genes responsible for salt tolerance.Funct Integr Genomics. 2019 Jan;19(1):61-73. doi: 10.1007/s10142-018-0628-6. Epub 2018 Jul 19. Funct Integr Genomics. 2019. PMID: 30046943
-
The salt-responsive transcriptome of chickpea roots and nodules via deepSuperSAGE.BMC Plant Biol. 2011 Feb 14;11:31. doi: 10.1186/1471-2229-11-31. BMC Plant Biol. 2011. PMID: 21320317 Free PMC article.
-
[Progress in molecular biology of a semi-mangrove, Millettia pinnata].Sheng Wu Gong Cheng Xue Bao. 2015 Apr;31(4):461-8. Sheng Wu Gong Cheng Xue Bao. 2015. PMID: 26380403 Review. Chinese.
-
Halophytes: Potential Resources for Salt Stress Tolerance Genes and Promoters.Front Plant Sci. 2017 May 18;8:829. doi: 10.3389/fpls.2017.00829. eCollection 2017. Front Plant Sci. 2017. PMID: 28572812 Free PMC article. Review.
Cited by
-
Analysis of stress-responsive transcriptome in the intestine of Asian seabass (Lates calcarifer) using RNA-seq.DNA Res. 2013 Oct;20(5):449-60. doi: 10.1093/dnares/dst022. Epub 2013 Jun 10. DNA Res. 2013. PMID: 23761194 Free PMC article.
-
Unravelling the MicroRNA-Mediated Gene Regulation in Developing Pongamia Seeds by High-Throughput Small RNA Profiling.Int J Mol Sci. 2019 Jul 17;20(14):3509. doi: 10.3390/ijms20143509. Int J Mol Sci. 2019. PMID: 31319494 Free PMC article.
-
The role of symbiotic nitrogen fixation in sustainable production of biofuels.Int J Mol Sci. 2014 Apr 29;15(5):7380-97. doi: 10.3390/ijms15057380. Int J Mol Sci. 2014. PMID: 24786096 Free PMC article.
-
Development and characterization of simple sequence repeat markers providing genome-wide coverage and high resolution in maize.DNA Res. 2013 Oct;20(5):497-509. doi: 10.1093/dnares/dst026. Epub 2013 Jun 26. DNA Res. 2013. PMID: 23804557 Free PMC article.
-
Comparative transcriptome analysis unveiling reactive oxygen species scavenging system of Sonneratia caseolaris under salinity stress.Front Plant Sci. 2022 Jul 25;13:953450. doi: 10.3389/fpls.2022.953450. eCollection 2022. Front Plant Sci. 2022. PMID: 35958196 Free PMC article.
References
-
- Flowers T.J. Improving crop salt tolerance. J. Exp. Bot. 2004;55:307–19. doi:10.1093/jxb/erh003. - DOI - PubMed
-
- Hasegawa P.M., Bressan R.A., Zhu J.K., Bohnert H. J. Plant cellular and molecular responses to high salinity. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2000;51:463–99. doi:10.1146/annurev.arplant.51.1.463. - DOI - PubMed
-
- Zhu J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002;53:247–73. doi:10.1146/annurev.arplant.53.091401.143329. - DOI - PMC - PubMed
-
- Seki M., Narusaka M., Ishida J., et al. Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J. 2002;31:279–92. doi:10.1046/j.1365-313X.2002.01359.x. - DOI - PubMed
-
- Jiang Y.Q., Deyholos M.K. Comprehensive transcriptional profiling of NaCl-stressed Arabidopsis roots reveals novel classes of responsive genes. BMC Plant Biol. 2006;6:25. doi:10.1186/1471-2229-6-25. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

