Expression of an entire bacterial operon in plants
- PMID: 22353575
- PMCID: PMC3320193
- DOI: 10.1104/pp.111.186197
Expression of an entire bacterial operon in plants
Abstract
Multigene expression is required for metabolic engineering, i.e. coregulated expression of all genes in a metabolic pathway for the production of a desired secondary metabolite. To that end, several transgenic approaches have been attempted with limited success. Better success has been achieved by transforming plastids with operons. IL-60 is a platform of constructs driven from the geminivirus Tomato yellow leaf curl virus. We demonstrate that IL-60 enables nontransgenic expression of an entire bacterial operon in tomato (Solanum lycopersicum) plants without the need for plastid (or any other) transformation. Delivery to the plant is simple, and the rate of expressing plants is close to 100%, eliminating the need for selectable markers. Using this platform, we show the expression of an entire metabolic pathway in plants and delivery of the end product secondary metabolite (pyrrolnitrin). Expression of this unique secondary metabolite resulted in the appearance of a unique plant phenotype disease resistance. Pyrrolnitrin production was already evident 2 d after application of the operon to plants and persisted throughout the plant's life span. Expression of entire metabolic pathways in plants is potentially beneficial for plant improvement, disease resistance, and biotechnological advances, such as commercial production of desired metabolites.
Figures






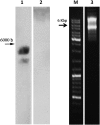
References
-
- Amoutzias G, Van de Peer Y. (2008) Together we stand: genes cluster to coordinate regulation. Dev Cell 14: 640–642 - PubMed
-
- Arima K, Imanaki H, Kousaka M, Fukuta A, Tamura G. (1964) Pyrrolnitrin, a new antibiotic substance, produced by Pseudomonas. Agric Biol Chem 28: 575–576
-
- Baily JA. (1982) Mechanisms of phytoalexine accumulation. In Mansfield, ed, Phytoalexins. Blackie, Glasgow, pp 289–318
-
- Blumenthal T. (2004) Operons in eukaryotes. Brief Funct Genomics Proteomics 3: 199–211 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

