Genome-Wide Scale-Free Network Inference for Candida albicans
- PMID: 22355294
- PMCID: PMC3280432
- DOI: 10.3389/fmicb.2012.00051
Genome-Wide Scale-Free Network Inference for Candida albicans
Abstract
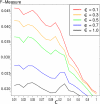
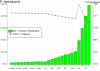
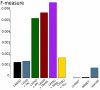
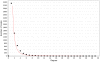
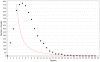
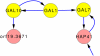
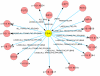
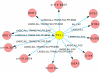
Discovery of essential genes in pathogenic organisms is an important step in the development of new medication. Despite a growing number of genome data available, little is known about C. albicans, a major fungal pathogen. Most of the human population carries C. albicans as commensal, but it can cause systemic infection that may lead to the death of the host if the immune system has deteriorated. In many organisms central nodes in the interaction network (hubs) play a crucial role for information and energy transport. Knock-outs of such hubs often lead to lethal phenotypes making them interesting drug targets. To identify these central genes via topological analysis, we inferred gene regulatory networks that are sparse and scale-free. We collected information from various sources to complement the limited expression data available. We utilized a linear regression algorithm to infer genome-wide gene regulatory interaction networks. To evaluate the predictive power of our approach, we used an automated text-mining system that scanned full-text research papers for known interactions. With the help of the compendium of known interactions, we also optimize the influence of the prior knowledge and the sparseness of the model to achieve the best results. We compare the results of our approach with those of other state-of-the-art network inference methods and show that we outperform those methods. Finally we identify a number of hubs in the genome of the fungus and investigate their biological relevance.
Keywords: Candida albicans; LASSO; hubs; linear regression; network inference; prior knowledge; reverse engineering; scale-free.
Figures

References
-
- Arnaud M. B., Costanzo M. C., Skrzypek M. S., Shah P., Binkley G., Lane C., Miyasato S. R., G S. (2010). Candida Genome Database. Available at: http://www.candidagenome.org/ - PMC - PubMed
-
- Buyko E., Faessler E., Wermter J., Hahn U. (2011). Syntactic simplification and semantic enrichment – trimming dependency graphs for event extraction. Comput. Intell. 27, 610–644 10.1111/j.1467-8640.2011.00402.x - DOI
-
- D’Enfert C., Hube B. (2007). Candida: Comparative and Functional Genomics. Norfolk: Caister Academic Press
LinkOut - more resources
Full Text Sources

