Uneven spread of cis- and trans-editing aminoacyl-tRNA synthetase domains within translational compartments of P. falciparum
- PMID: 22355703
- PMCID: PMC3240968
- DOI: 10.1038/srep00188
Uneven spread of cis- and trans-editing aminoacyl-tRNA synthetase domains within translational compartments of P. falciparum
Abstract
Accuracy of aminoacylation is dependent on maintaining fidelity during attachment of amino acids to cognate tRNAs. Cis- and trans-editing protein factors impose quality control during protein translation, and 8 of 36 Plasmodium falciparum aminoacyl-tRNA synthetase (aaRS) assemblies contain canonical putative editing modules. Based on expression and localization profiles of these 8 aaRSs, we propose an asymmetric distribution between the parasite cytoplasm and its apicoplast of putative editing-domain containing aaRSs. We also show that the single copy alanyl- and threonyl-tRNA synthetases are dually targeted to parasite cytoplasm and apicoplast. This bipolar presence of two unique synthetases presents opportunity for inhibitor targeting their aminoacylation and editing activities in twin parasite compartments. We used this approach to identify specific inhibitors against the alanyl- and threonyl-tRNA synthetases. Further development of such inhibitors may lead to anti-parasitics which simultaneously block protein translation in two key parasite organelles, a strategy of wider applicability for pathogen control.
Figures


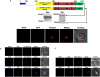


Similar articles
-
Plasmodium falciparum mitochondria import tRNAs along with an active phenylalanyl-tRNA synthetase.Biochem J. 2015 Feb 1;465(3):459-69. doi: 10.1042/BJ20140998. Biochem J. 2015. PMID: 25391660
-
A continuous assay for monitoring the synthetic and proofreading activities of multiple aminoacyl-tRNA synthetases for high-throughput drug discovery.RNA Biol. 2018;15(4-5):659-666. doi: 10.1080/15476286.2017.1397262. Epub 2017 Dec 15. RNA Biol. 2018. PMID: 29168435 Free PMC article.
-
A genomic glimpse of aminoacyl-tRNA synthetases in malaria parasite Plasmodium falciparum.BMC Genomics. 2009 Dec 31;10:644. doi: 10.1186/1471-2164-10-644. BMC Genomics. 2009. PMID: 20042123 Free PMC article.
-
Structural analyses of the malaria parasite aminoacyl-tRNA synthetases provide new avenues for antimalarial drug discovery.Protein Sci. 2021 Sep;30(9):1793-1803. doi: 10.1002/pro.4148. Protein Sci. 2021. PMID: 34184352 Free PMC article. Review.
-
Trans-editing by aminoacyl-tRNA synthetase-like editing domains.Enzymes. 2020;48:69-115. doi: 10.1016/bs.enz.2020.07.002. Epub 2020 Sep 8. Enzymes. 2020. PMID: 33837712 Review.
Cited by
-
Conformational heterogeneity in apo and drug-bound structures of Toxoplasma gondii prolyl-tRNA synthetase.Acta Crystallogr F Struct Biol Commun. 2019 Nov 1;75(Pt 11):714-724. doi: 10.1107/S2053230X19014808. Epub 2019 Nov 7. Acta Crystallogr F Struct Biol Commun. 2019. PMID: 31702585 Free PMC article.
-
Structural and functional analysis of the anti-malarial drug target prolyl-tRNA synthetase.J Struct Funct Genomics. 2014 Dec;15(4):181-90. doi: 10.1007/s10969-014-9186-x. Epub 2014 Jul 22. J Struct Funct Genomics. 2014. PMID: 25047712
-
Validation of Putative Apicoplast-Targeting Drugs Using a Chemical Supplementation Assay in Cultured Human Malaria Parasites.Antimicrob Agents Chemother. 2017 Dec 21;62(1):e01161-17. doi: 10.1128/AAC.01161-17. Print 2018 Jan. Antimicrob Agents Chemother. 2017. PMID: 29109165 Free PMC article.
-
Recent advances in the biology and drug targeting of malaria parasite aminoacyl-tRNA synthetases.Malar J. 2016 Apr 12;15:203. doi: 10.1186/s12936-016-1247-0. Malar J. 2016. PMID: 27068331 Free PMC article. Review.
-
Evolutionary and structural annotation of disease-associated mutations in human aminoacyl-tRNA synthetases.BMC Genomics. 2014 Dec 4;15(1):1063. doi: 10.1186/1471-2164-15-1063. BMC Genomics. 2014. PMID: 25476837 Free PMC article.
References
-
- WHO World malaria report 2009. Geneva, Switzerland<1/pub-tl> (2009).
-
- Miller L. H., Baruch D. I. Marsh K. & Doumbo O. K. The pathogenic basis of malaria. Int J. Parasitiol. 405, 673–679 (2002). - PubMed
-
- Ibba M. & Soll D. Aminoacyl-tRNA synthesis. Annu Rev Biochem. 69, 617–50 (2000). - PubMed
-
- Jakubowski H. Accuracy of aminoacyl-tRNA synthetases: proofreading of amino acids. In the Aminoacyl-tRNA Synthetases. ed. Ibba M., Francklyn, C. & Cusack, S. pp. 354–96 Austin, TX: LandesBiosci./Eurekah.com (2005).
-
- Ling J., Reynolds N. & Ibba M. Aminoacyl-tRNA synthesis and Translational quality control. Annu Rev Microbiol. 63, 61–78 (2009). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

