Phospho-ΔNp63α-dependent regulation of autophagic signaling through transcription and micro-RNA modulation
- PMID: 22356768
- PMCID: PMC3335921
- DOI: 10.4161/cc.11.6.19670
Phospho-ΔNp63α-dependent regulation of autophagic signaling through transcription and micro-RNA modulation
Abstract
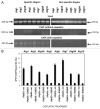
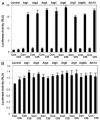
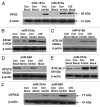
Cisplatin was shown to induce the ataxia telangiectasia mutated (ATM)-dependent phosphorylation of tumor protein p63 isoform, (ΔNp63α), leading to a transcriptional regulation of specific genes implicated in the control of cell death of squamous cell carcinoma (SCC) cells. We previously observed that the cisplatin-induced phosphorylated (p)-ΔNp63α transcriptionally regulates the expression of specific microRNAs (miRNAs) in SCC cells. We found here that cisplatin exposure of SCC cells led to modulation of the members of the autophagic pathway, such as Atg1/Ulk1, Atg3, Atg4A, Atg5, Atg6/Becn1, Atg7, Atg9A and Atg10, by a direct p-ΔNp63α-dependent transcriptional regulation. We further found that specific miRNAs (miR-181a, miR-519a, miR-374a and miR-630), which are critical downstream targets of the p-ΔNp63α, modulated the protein levels of ATG5, ATG6/BECN1, ATG10, ATG12, ATG16L1 and UVRAG, adding another level of expression control for autophagic pathways in SCC cells upon cisplatin exposure. Our data support the notion that the cisplatin-induced p-ΔNp63α could regulate key pathways implicated in response of cancer cells to chemotherapeutics.
Figures

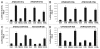
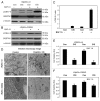
Similar articles
-
Phospho-ΔNp63α/SREBF1 protein interactions: bridging cell metabolism and cisplatin chemoresistance.Cell Cycle. 2012 Oct 15;11(20):3810-27. doi: 10.4161/cc.22022. Epub 2012 Sep 5. Cell Cycle. 2012. PMID: 22951905 Free PMC article.
-
Phospho-ΔNp63α/Rpn13-dependent regulation of LKB1 degradation modulates autophagy in cancer cells.Aging (Albany NY). 2010 Dec;2(12):959-68. doi: 10.18632/aging.100249. Aging (Albany NY). 2010. PMID: 21191146 Free PMC article.
-
Phospho-DeltaNp63alpha/NF-Y protein complex transcriptionally regulates DDIT3 expression in squamous cell carcinoma cells upon cisplatin exposure.Cell Cycle. 2010 Jan 15;9(2):328-38. doi: 10.4161/cc.9.2.10432. Epub 2010 Jan 26. Cell Cycle. 2010. PMID: 20023394
-
The guardians of the genome (p53, TA-p73, and TA-p63) are regulators of tumor suppressor miRNAs network.Cancer Metastasis Rev. 2010 Dec;29(4):613-39. doi: 10.1007/s10555-010-9257-9. Cancer Metastasis Rev. 2010. PMID: 20922462 Review.
-
TP63 links chromatin remodeling and enhancer reprogramming to epidermal differentiation and squamous cell carcinoma development.Cell Mol Life Sci. 2020 Nov;77(21):4325-4346. doi: 10.1007/s00018-020-03539-2. Epub 2020 May 23. Cell Mol Life Sci. 2020. PMID: 32447427 Free PMC article. Review.
Cited by
-
Tumor Protein (TP)-p53 Members as Regulators of Autophagy in Tumor Cells upon Marine Drug Exposure.Mar Drugs. 2016 Aug 16;14(8):154. doi: 10.3390/md14080154. Mar Drugs. 2016. PMID: 27537898 Free PMC article.
-
Autophagy-associated biomarkers ULK2, UVRAG, and miRNAs miR-21, miR-126, and miR-374: Prognostic significance in glioma patients.PLoS One. 2024 Sep 30;19(9):e0311308. doi: 10.1371/journal.pone.0311308. eCollection 2024. PLoS One. 2024. PMID: 39348350 Free PMC article.
-
Phospho-ΔNp63α/microRNA network modulates epigenetic regulatory enzymes in squamous cell carcinomas.Cell Cycle. 2014;13(5):749-61. doi: 10.4161/cc.27676. Epub 2014 Jan 6. Cell Cycle. 2014. PMID: 24394434 Free PMC article.
-
Human autophagy gene ATG16L1 is post-transcriptionally regulated by MIR142-3p.Autophagy. 2014 Mar;10(3):468-79. doi: 10.4161/auto.27553. Epub 2014 Jan 6. Autophagy. 2014. PMID: 24401604 Free PMC article.
-
Screening for E3-ubiquitin ligase inhibitors: challenges and opportunities.Oncotarget. 2014 Sep 30;5(18):7988-8013. doi: 10.18632/oncotarget.2431. Oncotarget. 2014. PMID: 25237759 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
