Function of homo- and hetero-oligomers of human nucleoplasmin/nucleophosmin family proteins NPM1, NPM2 and NPM3 during sperm chromatin remodeling
- PMID: 22362753
- PMCID: PMC3367197
- DOI: 10.1093/nar/gks162
Function of homo- and hetero-oligomers of human nucleoplasmin/nucleophosmin family proteins NPM1, NPM2 and NPM3 during sperm chromatin remodeling
Abstract
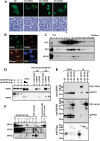
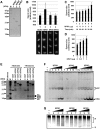
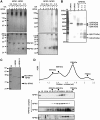
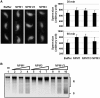
Sperm chromatin remodeling after oocyte entry is the essential step that initiates embryogenesis. This reaction involves the removal of sperm-specific basic proteins and chromatin assembly with histones. In mammals, three nucleoplasmin/nucleophosmin (NPM) family proteins-NPM1, NPM2 and NPM3-expressed in oocytes are presumed to cooperatively regulate sperm chromatin remodeling. We characterized the sperm chromatin decondensation and nucleosome assembly activities of three human NPM proteins. NPM1 and NPM2 mediated nucleosome assembly independently of other NPM proteins, whereas the function of NPM3 was largely dependent on formation of a complex with NPM1. Maximal sperm chromatin remodeling activity of NPM2 required the inhibition of its non-specific nucleic acid-binding activity by phosphorylation. Furthermore, the oligomer formation with NPM1 elicited NPM3 nucleosome assembly and sperm chromatin decondensation activity. NPM3 also suppressed the RNA-binding activity of NPM1, which enhanced the nucleoplasm-nucleolus shuttling of NPM1 in somatic cell nuclei. Our results proposed a novel mechanism whereby three NPM proteins cooperatively regulate chromatin disassembly and assembly in the early embryo and in somatic cells.
Figures




Similar articles
-
Drosophila TAP/p32 is a core histone chaperone that cooperates with NAP-1, NLP, and nucleophosmin in sperm chromatin remodeling during fertilization.Genes Dev. 2014 Sep 15;28(18):2027-40. doi: 10.1101/gad.248583.114. Genes Dev. 2014. PMID: 25228646 Free PMC article.
-
New insights into the nucleophosmin/nucleoplasmin family of nuclear chaperones.Bioessays. 2007 Jan;29(1):49-59. doi: 10.1002/bies.20512. Bioessays. 2007. PMID: 17187372 Review.
-
Vertebrate nucleoplasmin and NASP: egg histone storage proteins with multiple chaperone activities.FASEB J. 2012 Dec;26(12):4788-804. doi: 10.1096/fj.12-216663. Epub 2012 Sep 11. FASEB J. 2012. PMID: 22968912 Review.
-
The characterization of amphibian nucleoplasmins yields new insight into their role in sperm chromatin remodeling.BMC Genomics. 2006 Apr 28;7:99. doi: 10.1186/1471-2164-7-99. BMC Genomics. 2006. PMID: 16646973 Free PMC article.
-
NPM3, a member of the nucleophosmin/nucleoplasmin family, enhances activator-dependent transcription.Biochemistry. 2010 Feb 23;49(7):1355-7. doi: 10.1021/bi9021632. Biochemistry. 2010. PMID: 20073534
Cited by
-
Nuclear reprogramming of sperm and somatic nuclei in eggs and oocytes.Reprod Med Biol. 2013;12(4):133-149. doi: 10.1007/s12522-013-0155-z. Epub 2013 Jun 4. Reprod Med Biol. 2013. PMID: 24273450 Free PMC article.
-
Structure-function studies of a nucleoplasmin isoform from Plasmodium falciparum.J Biol Chem. 2025 Apr;301(4):108379. doi: 10.1016/j.jbc.2025.108379. Epub 2025 Mar 4. J Biol Chem. 2025. PMID: 40049416 Free PMC article.
-
Double maternal-effect: duplicated nucleoplasmin 2 genes, npm2a and npm2b, with essential but distinct functions are shared by fish and tetrapods.BMC Evol Biol. 2018 Nov 12;18(1):167. doi: 10.1186/s12862-018-1281-3. BMC Evol Biol. 2018. PMID: 30419815 Free PMC article.
-
Reprogramming the genome to totipotency in mouse embryos.Trends Cell Biol. 2015 Feb;25(2):82-91. doi: 10.1016/j.tcb.2014.09.006. Epub 2014 Oct 21. Trends Cell Biol. 2015. PMID: 25448353 Free PMC article. Review.
-
Leukemia-Associated Nup214 Fusion Proteins Disturb the XPO1-Mediated Nuclear-Cytoplasmic Transport Pathway and Thereby the NF-κB Signaling Pathway.Mol Cell Biol. 2016 Jun 15;36(13):1820-35. doi: 10.1128/MCB.00158-16. Print 2016 Jul 1. Mol Cell Biol. 2016. PMID: 27114368 Free PMC article.
References
-
- Laskey RA, Honda BM, Mills AD, Finch JT. Nucleosomes are assembled by an acidic protein which binds histones and transfers them to DNA. Nature. 1978;275:416–420. - PubMed
-
- Philpott A, Leno GH. Nucleoplasmin remodels sperm chromatin in Xenopus egg extracts. Cell. 1992;69:759–767. - PubMed
-
- Sylvestre EL, Pennetier S, Bureau M, Robert C, Sirard MA. Investigating the potential of genes preferentially expressed in oocyte to induce chromatin remodeling in somatic cells. Cell Reprogram. 2010;12:519–528. - PubMed
-
- Burns KH, Viveiros MM, Ren Y, Wang P, DeMayo FJ, Frail DE, Eppig JJ, Matzuk MM. Roles of NPM2 in chromatin and nucleolar organization in oocytes and embryos. Science. 2003;300:633–636. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

