Sample reproducibility of genetic association using different multimarker TDTs in genome-wide association studies: characterization and a new approach
- PMID: 22363405
- PMCID: PMC3281822
- DOI: 10.1371/journal.pone.0029613
Sample reproducibility of genetic association using different multimarker TDTs in genome-wide association studies: characterization and a new approach
Abstract
Multimarker Transmission/Disequilibrium Tests (TDTs) are very robust association tests to population admixture and structure which may be used to identify susceptibility loci in genome-wide association studies. Multimarker TDTs using several markers may increase power by capturing high-degree associations. However, there is also a risk of spurious associations and power reduction due to the increase in degrees of freedom. In this study we show that associations found by tests built on simple null hypotheses are highly reproducible in a second independent data set regardless the number of markers. As a test exhibiting this feature to its maximum, we introduce the multimarker 2-Groups TDT (mTDT(2G)), a test which under the hypothesis of no linkage, asymptotically follows a χ2 distribution with 1 degree of freedom regardless the number of markers. The statistic requires the division of parental haplotypes into two groups: disease susceptibility and disease protective haplotype groups. We assessed the test behavior by performing an extensive simulation study as well as a real-data study using several data sets of two complex diseases. We show that mTDT(2G) test is highly efficient and it achieves the highest power among all the tests used, even when the null hypothesis is tested in a second independent data set. Therefore, mTDT(2G) turns out to be a very promising multimarker TDT to perform genome-wide searches for disease susceptibility loci that may be used as a preprocessing step in the construction of more accurate genetic models to predict individual susceptibility to complex diseases.
Conflict of interest statement
Figures

 (left plot),
(left plot),  (plot in the middle) and
(plot in the middle) and  (right plot). A nominal level of
(right plot). A nominal level of  and a relative risk of
and a relative risk of  were used for all plots. Results for
were used for all plots. Results for  and
and  are plotted in purple circles and blue triangles respectively. Dashed lines show results for the data subset (125 trios randomly chosen) used to build the model while solid lines show results for a second data subset (the remaining 125 trios) used to test reproducibility.
are plotted in purple circles and blue triangles respectively. Dashed lines show results for the data subset (125 trios randomly chosen) used to build the model while solid lines show results for a second data subset (the remaining 125 trios) used to test reproducibility.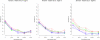
 family trios as a function of the recombination rate using the recessive and one-locus genetic model and haplotypes of lengths
family trios as a function of the recombination rate using the recessive and one-locus genetic model and haplotypes of lengths  (left plot),
(left plot),  (plot in the middle) and
(plot in the middle) and  (right plot). A nominal level of
(right plot). A nominal level of  and a relative risk of
and a relative risk of  were used for all plots. Results for
were used for all plots. Results for 

 and
and  i.e. all tests were applied under the holdout approach, are plotted in purple circles, blue triangles, green squares and red diamonds respectively. Dashed lines show results for a data subset of
i.e. all tests were applied under the holdout approach, are plotted in purple circles, blue triangles, green squares and red diamonds respectively. Dashed lines show results for a data subset of  trios randomly chosen while solid lines show results for a second data subset of
trios randomly chosen while solid lines show results for a second data subset of  trios used to test reproducibility of the holdout approach.
trios used to test reproducibility of the holdout approach.
 family trios as a function of the proportion of missing haplotypes using the additive and one-locus genetic model and haplotypes of lengths
family trios as a function of the proportion of missing haplotypes using the additive and one-locus genetic model and haplotypes of lengths  (left plot),
(left plot),  (plot in the middle) and
(plot in the middle) and  (right plot). A nominal level of
(right plot). A nominal level of  and a relative risk of
and a relative risk of  were used for all plots. Results for
were used for all plots. Results for 

 and
and  i.e. all tests were applied under the holdout approach, are plotted in purple circles, blue triangles, green squares and red diamonds respectively.
i.e. all tests were applied under the holdout approach, are plotted in purple circles, blue triangles, green squares and red diamonds respectively.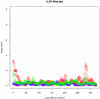
 TDTs used were
TDTs used were  (red diamonds),
(red diamonds),  (green squares),
(green squares),  (purple circles) and
(purple circles) and  (blue triangles).
(blue triangles).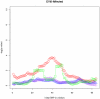
 TDTs used were
TDTs used were  (red diamonds),
(red diamonds),  (green squares),
(green squares),  (purple circles) and
(purple circles) and  (blue triangles).
(blue triangles).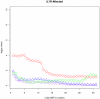
 TDTs used were
TDTs used were  (red diamonds),
(red diamonds),  (green squares),
(green squares),  (purple circles) and
(purple circles) and  (blue triangles).
(blue triangles).References
-
- BickeBöller H, Clerget-Darpoux F. Statistical properties of the allelic and genotypic transmission/disequilibrium test for multiallelic markers. Genetic Epidemiology. 1995;12:865–70. - PubMed
-
- Sham PC, Curtis D. An extended transmission/disequilibrium test (tdt) for multiallelic marker loci. Annals of Human Genetics. 1995;59:323–336. - PubMed
-
- Yu K, Gu CC, Xiong C, An P, Province M. Global Transmission/Disequilibrium tests based on haplotype sharing in multiple candidate genes. Genetic Epidemiology. 2005;29:223–35. - PubMed

