Microbial scout hypothesis, stochastic exit from dormancy, and the nature of slow growers
- PMID: 22367083
- PMCID: PMC3346438
- DOI: 10.1128/AEM.07307-11
Microbial scout hypothesis, stochastic exit from dormancy, and the nature of slow growers
Abstract
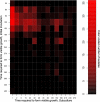
We recently proposed a scout model of the microbial life cycle (S. S. Epstein, Nature 457:1083, 2009), the central element of which is the hypothesis that dormant microbial cells wake up into active (so-called scout) cells stochastically, independently of environmental cues. Here, we check the principal prediction of this hypothesis: under growth-permissive conditions, dormant cells initiate growth at random time intervals and exhibit no species-specific lag phase. We show that a range of microorganisms, including environmental species, Escherichia coli, and Mycobacterium smegmatis, indeed wake up in a seemingly stochastic manner and independently of environmental conditions, even in the longest incubations conducted (months to years long). As is implicit in the model, most of the cultures we obtained after long incubations were not inherently slow growers. Of the environmental isolates that required ≥7 months to form visible growth, only 5% needed an equally long incubation upon subculturing, with the majority exhibiting regrowth within 24 to 48 h. This apparent change was not a result of adaptive mutation; rather, most microbial species that appear to be slow growers were in fact fast growers with a delayed initiation of division. Genuine slow growth thus appears to be less significant than previously believed. Random, low-frequency exit from the nongrowing state may be a key element of a general microbial survival strategy, and the phylogenetic breadth of the organisms exhibiting such exit indicates that it represents a general phenomenon. The stochasticity of awakening can also provide a parsimonious explanation to several microbiological observations, including the apparent randomness of latent infections and the existence of viable-but-nonculturable cells (VBNC).
Figures


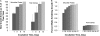
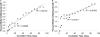
Similar articles
-
Viable but Nonculturable and Persister Cells Coexist Stochastically and Are Induced by Human Serum.Infect Immun. 2015 Nov;83(11):4194-203. doi: 10.1128/IAI.00404-15. Epub 2015 Aug 17. Infect Immun. 2015. PMID: 26283335 Free PMC article.
-
The role of histone-like protein, Hlp, in Mycobacterium smegmatis dormancy.FEMS Microbiol Lett. 2010 Jul;308(2):101-7. doi: 10.1111/j.1574-6968.2010.01988.x. Epub 2010 Apr 14. FEMS Microbiol Lett. 2010. PMID: 20497227
-
Characterization of a Mycobacterium smegmatis uvrA mutant impaired in dormancy induced by hypoxia and low carbon concentration.BMC Microbiol. 2011 Oct 18;11:231. doi: 10.1186/1471-2180-11-231. BMC Microbiol. 2011. PMID: 22008214 Free PMC article.
-
The "comfort timing" strategy: a potential pathway for the cultivation of uncultured microorganisms and a possible adaptation for environmental colonisation.FEMS Microbiol Ecol. 2023 Mar 23;99(4):fiad026. doi: 10.1093/femsec/fiad026. FEMS Microbiol Ecol. 2023. PMID: 36921985 Review.
-
Significance of Viable but Nonculturable Escherichia coli: Induction, Detection, and Control.J Microbiol Biotechnol. 2017 Mar 28;27(3):417-428. doi: 10.4014/jmb.1609.09063. J Microbiol Biotechnol. 2017. PMID: 27974738 Review.
Cited by
-
Soil microbe active community composition and capability of responding to litter addition after 12 years of no inputs.Appl Environ Microbiol. 2013 Feb;79(4):1385-92. doi: 10.1128/AEM.03181-12. Epub 2012 Dec 21. Appl Environ Microbiol. 2013. PMID: 23263952 Free PMC article.
-
Divergent extremes but convergent recovery of bacterial and archaeal soil communities to an ongoing subterranean coal mine fire.ISME J. 2017 Jun;11(6):1447-1459. doi: 10.1038/ismej.2017.1. Epub 2017 Mar 10. ISME J. 2017. PMID: 28282042 Free PMC article.
-
Constant Flux of Spatial Niche Partitioning through High-Resolution Sampling of Magnetotactic Bacteria.Appl Environ Microbiol. 2017 Sep 29;83(20):e01382-17. doi: 10.1128/AEM.01382-17. Print 2017 Oct 15. Appl Environ Microbiol. 2017. PMID: 28778897 Free PMC article.
-
Mycobacterial Growth.Cold Spring Harb Perspect Med. 2015 May 8;5(10):a021097. doi: 10.1101/cshperspect.a021097. Cold Spring Harb Perspect Med. 2015. PMID: 25957314 Free PMC article. Review.
-
Principles of seed banks and the emergence of complexity from dormancy.Nat Commun. 2021 Aug 10;12(1):4807. doi: 10.1038/s41467-021-24733-1. Nat Commun. 2021. PMID: 34376641 Free PMC article. Review.
References
-
- Avery SV. 2006. Microbial cell individuality and the underlying sources of heterogeneity. Nat. Rev. Microbiol. 4:577–587 - PubMed
-
- Baker GC, Smith JJ, Cowan DA. 2003. Review and re-analysis of domain-specific 16S primers. J. Microbiol. Methods 55:541–555 - PubMed
-
- Balaban NQ, Merrin J, Chait R, Kowalik L, Leibler S. 2004. Bacterial persistence as a phenotypic switch. Science 305:1622–1625 - PubMed
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

