Resampling procedures to identify important SNPs using a consensus approach
- PMID: 22373247
- PMCID: PMC3287897
- DOI: 10.1186/1753-6561-5-S9-S59
Resampling procedures to identify important SNPs using a consensus approach
Abstract
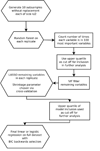
Our goal is to identify common single-nucleotide polymorphisms (SNPs) (minor allele frequency > 1%) that add predictive accuracy above that gained by knowledge of easily measured clinical variables. We take an algorithmic approach to predict each phenotypic variable using a combination of phenotypic and genotypic predictors. We perform our procedure on the first simulated replicate and then validate against the others. Our procedure performs well when predicting Q1 but is less successful for the other outcomes. We use resampling procedures where possible to guard against false positives and to improve generalizability. The approach is based on finding a consensus regarding important SNPs by applying random forests and the least absolute shrinkage and selection operator (LASSO) on multiple subsamples. Random forests are used first to discard unimportant predictors, narrowing our focus to roughly 100 important SNPs. A cross-validation LASSO is then used to further select variables. We combine these procedures to guarantee that cross-validation can be used to choose a shrinkage parameter for the LASSO. If the clinical variables were unavailable, this prefiltering step would be essential. We perform the SNP-based analyses simultaneously rather than one at a time to estimate SNP effects in the presence of other causal variants. We analyzed the first simulated replicate of Genetic Analysis Workshop 17 without knowledge of the true model. Post-conference knowledge of the simulation parameters allowed us to investigate the limitations of our approach. We found that many of the false positives we identified were substantially correlated with genuine causal SNPs.
Figures
References
-
- Breiman L. Random forests. Machine Learning. 2001;45:5–32. doi: 10.1023/A:1010933404324. - DOI
-
- Breiman L. Statistical modeling: the two cultures. Stat Sci. 2001;16:199–231. (with comments and a rejoinder by the author)
LinkOut - more resources
Full Text Sources

