Dynamic modelling under uncertainty: the case of Trypanosoma brucei energy metabolism
- PMID: 22379410
- PMCID: PMC3269904
- DOI: 10.1371/journal.pcbi.1002352
Dynamic modelling under uncertainty: the case of Trypanosoma brucei energy metabolism
Abstract
Kinetic models of metabolism require detailed knowledge of kinetic parameters. However, due to measurement errors or lack of data this knowledge is often uncertain. The model of glycolysis in the parasitic protozoan Trypanosoma brucei is a particularly well analysed example of a quantitative metabolic model, but so far it has been studied with a fixed set of parameters only. Here we evaluate the effect of parameter uncertainty. In order to define probability distributions for each parameter, information about the experimental sources and confidence intervals for all parameters were collected. We created a wiki-based website dedicated to the detailed documentation of this information: the SilicoTryp wiki (http://silicotryp.ibls.gla.ac.uk/wiki/Glycolysis). Using information collected in the wiki, we then assigned probability distributions to all parameters of the model. This allowed us to sample sets of alternative models, accurately representing our degree of uncertainty. Some properties of the model, such as the repartition of the glycolytic flux between the glycerol and pyruvate producing branches, are robust to these uncertainties. However, our analysis also allowed us to identify fragilities of the model leading to the accumulation of 3-phosphoglycerate and/or pyruvate. The analysis of the control coefficients revealed the importance of taking into account the uncertainties about the parameters, as the ranking of the reactions can be greatly affected. This work will now form the basis for a comprehensive Bayesian analysis and extension of the model considering alternative topologies.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
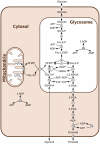

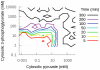
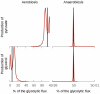
 parameter to 0). In aerobic conditions the effect is more important, indicating that this division is not entirely due to the topology of the model in this case.
parameter to 0). In aerobic conditions the effect is more important, indicating that this division is not entirely due to the topology of the model in this case.
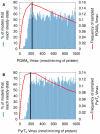
 (B) Percentage of models that reach steady-state within 300 minutes as a function of pyruvate transport
(B) Percentage of models that reach steady-state within 300 minutes as a function of pyruvate transport  . The red line is the distribution of the parameter as it is usually sampled. The black line is the fixed-parameter value. A model which has a value for one of these two parameters smaller than the mean will easily fail to reach steady-state, whatever the other parameter values and despite these
. The red line is the distribution of the parameter as it is usually sampled. The black line is the fixed-parameter value. A model which has a value for one of these two parameters smaller than the mean will easily fail to reach steady-state, whatever the other parameter values and despite these  values still being close to their mean. This reveals fragilities in the model.
values still being close to their mean. This reveals fragilities in the model.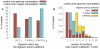

 values retrieved from the database (green) and a log-normal distribution with the same mean and standard-deviation (red) are shown.
values retrieved from the database (green) and a log-normal distribution with the same mean and standard-deviation (red) are shown.References
-
- Borger S, Liebermeister W, Klipp E. Prediction of enzyme kinetic parameters based on statistical learning. Genome Inform Ser. 2006;17:80–87. - PubMed
-
- Rizk ML, Liao JC. Ensemble modeling and related mathematical modeling of metabolic networks. J Taiwan Inst Chem E. 2009;40:595–601.
-
- Mišković L, Hatzimanikatis V. Modeling of uncertainties in biochemical reactions. Biotechnol Bioeng. 2011;108:413–423. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

