Single enantiomer of YK-4-279 demonstrates specificity in targeting the oncogene EWS-FLI1
- PMID: 22383402
- PMCID: PMC3326647
- DOI: 10.18632/oncotarget.454
Single enantiomer of YK-4-279 demonstrates specificity in targeting the oncogene EWS-FLI1
Abstract
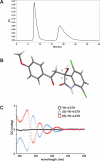
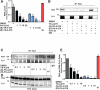
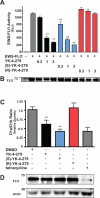
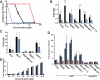
Oncogenic fusion proteins, such as EWS-FLI1, are excellent therapeutic targets as they are only located within the tumor. However, there are currently no agents targeted toward transcription factors, which are often considered to be 'undruggable.' A considerable body of evidence is accruing that refutes this claim based upon the intrinsic disorder of transcription factors. Our previous studies show that RNA Helicase A (RHA) enhances the oncogenesis of EWS-FLI1, a putative intrinsically disordered protein. Interruption of this protein-protein complex by small molecule inhibitors validates this interaction as a unique therapeutic target. Single enantiomer activity from a chiral compound has been recognized as strong evidence for specificity in a small molecule-protein interaction. Our compound, YK-4-279, has a chiral center and can be separated into two enantiomers by chiral HPLC. We show that there is a significant difference in activity between the two enantiomers. (S)-YK-4-279 is able to disrupt binding between EWS-FLI1 and RHA in an immunoprecipitation assay and blocks the transcriptional activity of EWS-FLI1, while (R)-YK-4-279 cannot. Enantiospecific effects are also established in cytotoxicity assays and caspase assays, where up to a log-fold difference is seen between (S)-YK-4-279 and the racemic YK-4-279. Our findings indicate that only one enantiomer of our small molecule is able to specifically target a protein-protein interaction. This work is significant for its identification of a single enantiomer effect upon a protein interaction suggesting that small molecule targeting of intrinsically disordered proteins can be specific. Furthermore, proving YK-4-279 has only one functional enantiomer will be helpful in moving this compound towards clinical trials.
Figures
References
-
- Kim J, Pelletier J. Molecular genetics of chromosome translocations involving EWS and related family members. Physiol Genomics. 1999;1:127–138. - PubMed
-
- Delattre O, Zucman J, Plougastel B, Desmaze C, Melot T, Peter M, Kovar H, Joubert I, de Jong P, Rouleau G, et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature. 1992;359:162–165. - PubMed
-
- Darnell JE., Jr Transcription factors as targets for cancer therapy. Nature reviews Cancer. 2002;2:740–749. - PubMed
-
- Wright PE, Dyson HJ. Intrinsically unstructured proteins: re-assessing the protein structure-function paradigm. Journal of molecular biology. 1999;293:321–331. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

