Human mitochondrial DNA helicase TWINKLE is both an unwinding and annealing helicase
- PMID: 22383523
- PMCID: PMC3340288
- DOI: 10.1074/jbc.M111.309468
Human mitochondrial DNA helicase TWINKLE is both an unwinding and annealing helicase
Abstract
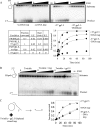
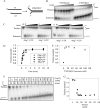
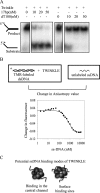
TWINKLE is a nucleus-encoded human mitochondrial (mt)DNA helicase. Point mutations in TWINKLE are associated with heritable neuromuscular diseases characterized by deletions in the mtDNA. To understand the biochemical basis of these diseases, it is important to define the roles of TWINKLE in mtDNA metabolism by studying its enzymatic activities. To this end, we purified native TWINKLE from Escherichia coli. The recombinant TWINKLE assembles into hexamers and higher oligomers, and addition of MgUTP stabilizes hexamers over higher oligomers. Probing into the DNA unwinding activity, we discovered that the efficiency of unwinding is greatly enhanced in the presence of a heterologous single strand-binding protein or a single-stranded (ss) DNA that is complementary to the unwound strand. We show that TWINKLE, although a helicase, has an antagonistic activity of annealing two complementary ssDNAs that interferes with unwinding in the absence of gp2.5 or ssDNA trap. Furthermore, only ssDNA and not double-stranded (ds)DNA competitively inhibits the annealing activity, although both DNAs bind with high affinities. This implies that dsDNA binds to a site that is distinct from the ssDNA-binding site that promotes annealing. Fluorescence anisotropy competition binding experiments suggest that TWINKLE has more than one ssDNA-binding sites, and we speculate that a surface-exposed ssDNA-specific site is involved in catalyzing DNA annealing. We propose that the strand annealing activity of TWINKLE may play a role in recombination-mediated replication initiation found in the mitochondria of mammalian brain and heart or in replication fork regression during repair of damaged DNA replication forks.
Figures

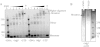
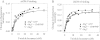

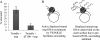
Similar articles
-
Stimulation of Variant Forms of the Mitochondrial DNA Helicase Twinkle by the Mitochondrial Single-Stranded DNA-Binding Protein.Methods Mol Biol. 2021;2281:313-322. doi: 10.1007/978-1-0716-1290-3_20. Methods Mol Biol. 2021. PMID: 33847968
-
Homologous DNA strand exchange activity of the human mitochondrial DNA helicase TWINKLE.Nucleic Acids Res. 2016 May 19;44(9):4200-10. doi: 10.1093/nar/gkw098. Epub 2016 Feb 16. Nucleic Acids Res. 2016. PMID: 26887820 Free PMC article.
-
Single-molecule level structural dynamics of DNA unwinding by human mitochondrial Twinkle helicase.J Biol Chem. 2020 Apr 24;295(17):5564-5576. doi: 10.1074/jbc.RA120.012795. Epub 2020 Mar 25. J Biol Chem. 2020. PMID: 32213598 Free PMC article.
-
TWINKLE and Other Human Mitochondrial DNA Helicases: Structure, Function and Disease.Genes (Basel). 2020 Apr 9;11(4):408. doi: 10.3390/genes11040408. Genes (Basel). 2020. PMID: 32283748 Free PMC article. Review.
-
Mechanistic and biological aspects of helicase action on damaged DNA.Cell Cycle. 2010 Jun 15;9(12):2317-29. doi: 10.4161/cc.9.12.11902. Epub 2010 Jun 15. Cell Cycle. 2010. PMID: 20574162 Free PMC article. Review.
Cited by
-
Structure, function and evolution of the animal mitochondrial replicative DNA helicase.Crit Rev Biochem Mol Biol. 2016;51(1):53-64. doi: 10.3109/10409238.2015.1117056. Epub 2015 Nov 29. Crit Rev Biochem Mol Biol. 2016. PMID: 26615986 Free PMC article. Review.
-
Biochemical Characterization of the Human Mitochondrial Replicative Twinkle Helicase: SUBSTRATE SPECIFICITY, DNA BRANCH MIGRATION, AND ABILITY TO OVERCOME BLOCKADES TO DNA UNWINDING.J Biol Chem. 2016 Jul 1;291(27):14324-14339. doi: 10.1074/jbc.M115.712026. Epub 2016 May 11. J Biol Chem. 2016. PMID: 27226550 Free PMC article.
-
Autoregulation of the real-time kinetics of the human mitochondrial replicative helicase.Nat Commun. 2025 Jul 1;16(1):5460. doi: 10.1038/s41467-025-60289-0. Nat Commun. 2025. PMID: 40592829 Free PMC article.
-
Twinkle mutations in two Chinese families with autosomal dominant progressive external ophthalmoplegia.Neurol Sci. 2014 Mar;35(3):443-8. doi: 10.1007/s10072-013-1557-8. Epub 2013 Oct 4. Neurol Sci. 2014. PMID: 24091712
-
DNA helicases associated with genetic instability, cancer, and aging.Adv Exp Med Biol. 2013;767:123-44. doi: 10.1007/978-1-4614-5037-5_6. Adv Exp Med Biol. 2013. PMID: 23161009 Free PMC article.
References
-
- Spelbrink J. N., Li F. Y., Tiranti V., Nikali K., Yuan Q. P., Tariq M., Wanrooij S., Garrido N., Comi G., Morandi L., Santoro L., Toscano A., Fabrizi G. M., Somer H., Croxen R., Beeson D., Poulton J., Suomalainen A., Jacobs H. T., Zeviani M., Larsson C. (2001) Human mitochondrial DNA deletions associated with mutations in the gene encoding Twinkle, a phage T7 gene 4-like protein localized in mitochondria. Nat. Genet. 28, 223–231 - PubMed
-
- Korhonen J. A., Pande V., Holmlund T., Farge G., Pham X. H., Nilsson L., Falkenberg M. (2008) Structure-function defects of the TWINKLE linker region in progressive external ophthalmoplegia. J. Mol. Biol. 377, 691–705 - PubMed
-
- Patel S. S., Picha K. M. (2000) Structure and function of hexameric helicases. Annu. Rev. Biochem. 69, 651–697 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

