Frequent incorporation of ribonucleotides during HIV-1 reverse transcription and their attenuated repair in macrophages
- PMID: 22383524
- PMCID: PMC3340154
- DOI: 10.1074/jbc.M112.348482
Frequent incorporation of ribonucleotides during HIV-1 reverse transcription and their attenuated repair in macrophages
Abstract
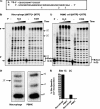
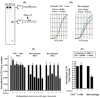
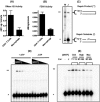
Macrophages are well known long-lived reservoirs of HIV-1. Unlike activated CD4(+) T cells, this nondividing HIV-1 target cell type contains a very low level of the deoxynucleoside triphosphates (dNTPs) required for proviral DNA synthesis whereas the ribonucleoside triphosphate (rNTP) levels remain in the millimolar range, resulting in an extremely low dNTP/rNTP ratio. Biochemical simulations demonstrate that HIV-1 reverse transcriptase (RT) efficiently incorporates ribonucleoside monophosphates (rNMPs) during DNA synthesis at this ratio, predicting frequent rNMP incorporation by the virus specifically in macrophages. Indeed, HIV-1 RT incorporates rNMPs at a remarkable rate of 1/146 nucleotides during macrophage infection. This greatly exceeds known rates for cellular replicative polymerases. In contrast, little or no rNMP incorporation is detected in CD4(+) T cells. Repair of these rNMP lesions is also substantially delayed in macrophages compared with CD4(+) T cells. Single rNMPs embedded in a DNA template are known to induce cellular DNA polymerase pausing, which mechanistically contributes to mutation synthesis. Indeed, we also observed that embedded rNMPs in a dsDNA template also induce HIV-1 RT DNA synthesis pausing. Moreover, unrepaired rNMPs incorporated into the provirus during HIV-1 reverse transcription would be generally mutagenic as was shown in Saccharomyces cerevisiae. Most importantly, the frequent incorporation of rNMPs makes them an ideal candidate for development of a new class of HIV RT inhibitors.
Figures


Similar articles
-
Viral protein X reduces the incorporation of mutagenic noncanonical rNTPs during lentivirus reverse transcription in macrophages.J Biol Chem. 2020 Jan 10;295(2):657-666. doi: 10.1074/jbc.RA119.011466. Epub 2019 Dec 5. J Biol Chem. 2020. PMID: 31806704 Free PMC article.
-
Pre-steady state kinetic analysis of HIV-1 reverse transcriptase for non-canonical ribonucleoside triphosphate incorporation and DNA synthesis from ribonucleoside-containing DNA template.Antiviral Res. 2015 Mar;115:75-82. doi: 10.1016/j.antiviral.2014.12.016. Epub 2014 Dec 31. Antiviral Res. 2015. PMID: 25557601 Free PMC article.
-
Effect of ribonucleotides embedded in a DNA template on HIV-1 reverse transcription kinetics and fidelity.J Biol Chem. 2013 May 3;288(18):12522-32. doi: 10.1074/jbc.M113.458398. Epub 2013 Mar 11. J Biol Chem. 2013. PMID: 23479739 Free PMC article.
-
RNases H1 and H2: guardians of the stability of the nuclear genome when supply of dNTPs is limiting for DNA synthesis.Curr Genet. 2020 Dec;66(6):1073-1084. doi: 10.1007/s00294-020-01086-8. Epub 2020 Sep 4. Curr Genet. 2020. PMID: 32886170 Free PMC article. Review.
-
RNase H2-RED carpets the path to eukaryotic RNase H2 functions.DNA Repair (Amst). 2019 Dec;84:102736. doi: 10.1016/j.dnarep.2019.102736. Epub 2019 Oct 23. DNA Repair (Amst). 2019. PMID: 31761672 Free PMC article. Review.
Cited by
-
Intracellular nucleotide levels and the control of retroviral infections.Virology. 2013 Feb 20;436(2):247-54. doi: 10.1016/j.virol.2012.11.010. Epub 2012 Dec 20. Virology. 2013. PMID: 23260109 Free PMC article. Review.
-
Impact of 1,N6-ethenoadenosine, a damaged ribonucleotide in DNA, on translesion synthesis and repair.J Biol Chem. 2020 May 1;295(18):6092-6107. doi: 10.1074/jbc.RA120.012829. Epub 2020 Mar 25. J Biol Chem. 2020. PMID: 32213600 Free PMC article.
-
Reduction of hRNase H2 activity in Aicardi-Goutières syndrome cells leads to replication stress and genome instability.Hum Mol Genet. 2015 Feb 1;24(3):649-58. doi: 10.1093/hmg/ddu485. Epub 2014 Sep 30. Hum Mol Genet. 2015. PMID: 25274781 Free PMC article.
-
Intertwined: SAMHD1 cellular functions, restriction, and viral evasion strategies.Med Microbiol Immunol. 2019 Aug;208(3-4):513-529. doi: 10.1007/s00430-019-00593-x. Epub 2019 Mar 16. Med Microbiol Immunol. 2019. PMID: 30879196 Review.
-
SAMHD1 Functions and Human Diseases.Viruses. 2020 Mar 31;12(4):382. doi: 10.3390/v12040382. Viruses. 2020. PMID: 32244340 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

