Fructose-bisphophate aldolase exhibits functional roles between carbon metabolism and the hrp system in rice pathogen Xanthomonas oryzae pv. oryzicola
- PMID: 22384086
- PMCID: PMC3285194
- DOI: 10.1371/journal.pone.0031855
Fructose-bisphophate aldolase exhibits functional roles between carbon metabolism and the hrp system in rice pathogen Xanthomonas oryzae pv. oryzicola
Abstract
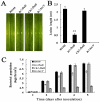
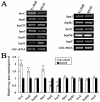
Fructose-bisphophate aldolase (FbaB), is an enzyme in glycolysis and gluconeogenesis in living organisms. The mutagenesis in a unique fbaB gene of Xanthomonas oryzae pv. oryzicola, the causal agent of rice bacterial leaf streak, led the pathogen not only unable to use pyruvate and malate for growth and delayed its growth when fructose was used as the sole carbon source, but also reduced extracellular polysaccharide (EPS) production and impaired bacterial virulence and growth in rice. Intriguingly, the fbaB promoter contains an imperfect PIP-box (plant-inducible promoter) (TTCGT-N(9)-TTCGT). The expression of fbaB was negatively regulated by a key hrp regulatory HrpG and HrpX cascade. Base substitution in the PIP-box altered the regulation of fbaB with the cascade. Furthermore, the expression of fbaB in X. oryzae pv. oryzicola RS105 strain was inducible in planta rather than in a nutrient-rich medium. Except other hrp-hrc-hpa genes, the expression of hrpG and hrpX was repressed and the transcripts of hrcC, hrpE and hpa3 were enhanced when fbaB was deleted. The mutation in hrcC, hrpE or hpa3 reduced the ability of the pathogen to acquire pyruvate and malate. In addition, bacterial virulence and growth in planta and EPS production in RΔfbaB mutant were completely restored to the wild-type level by the presence of fbaB in trans. This is the first report to demonstrate that carbohydrates, assimilated by X. oryzae pv. oryzicola, play critical roles in coordinating hrp gene expression through a yet unknown regulator.
Conflict of interest statement
Figures


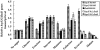
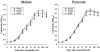
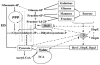
References
-
- Eisenreich W, Dandekar T, Heesemann J, Goebel W. Carbon metabolism of intracellular bacterial pathogens and possible links to virulence. Nat Rev Microbiol. 2010;8:401–412. - PubMed
-
- Wang LF, Rong W, He CZ. Two Xanthomonas extracellular polygalacturonases, PghAxc and PghBxc, are regulated by type III secretion regulators HrpX and HrpG and are required for virulence. Mol Plant-Microbe Interact. 2008;21:555–563. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

