Methylome of fetal and maternal monocytes and macrophages at the feto-maternal interface
- PMID: 22385097
- PMCID: PMC3479407
- DOI: 10.1111/j.1600-0897.2012.01108.x
Methylome of fetal and maternal monocytes and macrophages at the feto-maternal interface
Abstract
Problem: Decidual macrophages (dMφ) of the mother and placental macrophages (Hofbauer cells, HC) of the fetus are deployed at a critical location: the feto-maternal interface. This study was conducted to compare the DNA methylome of maternal and fetal monocytes, dMφ, and HC and thereby to determine the immunobiological importance of DNA methylation in pregnancy.
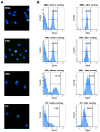
Method of study: Paired samples were obtained from normal pregnant women at term not in labor and their neonates. Maternal monocytes (MMo) and fetal monocytes (FMo) were isolated from the peripheral blood of mothers and fetal cord blood, respectively. dMφ and HC were obtained from the decidua of fetal membranes and placentas, respectively. DNA methylation profiling was performed using the Illumina Infinium Human Methylation27 BeadChip. Quantitative real-time PCR and Western Blot were performed for validation experiments.
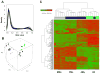
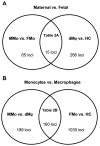
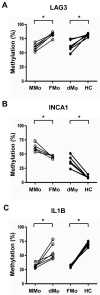
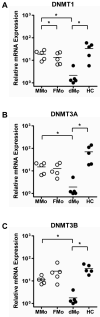
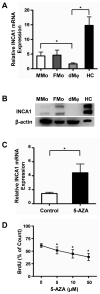
Results: (i) Significant differences in DNA methylation were found in each comparison (MMo versus FMo, 65 loci; dMφ versus HC, 266 loci; MMo versus dMφ, 199 loci; FMo versus HC, 1030 loci). (ii) Many of the immune response-related genes were hypermethylated in fetal cells (FMo and HC) compared to maternal cells (MMo and dMφ). (iii) Genes encoding markers of classical macrophage activation were hypermethylated, and genes encoding alternative macrophage activation were hypomethylated in dMφ and HC compared to MMo and FMo, respectively. (iv) mRNA expressions of DNMT1, DNMT3A, and DNMT3B were significantly lower in dMφ than in HC. (v) 5-azacytidine treatment increased expression of INCA1 in dMφ.
Conclusions: The findings herein indicate that DNA methylation patterns change during monocyte-macrophage differentiation at the feto-maternal interface. It is also suggested that DNA methylation is an important component of the biological machinery conferring an anti-inflammatory phenotype to macrophages at the feto-maternal interface.
© 2012 John Wiley & Sons A/S.
Conflict of interest statement
The authors have no financial conflicts of interest.
Figures
References
-
- Satosar A, Ramirez NC, Bartholomew D, Davis J, Nuovo GJ. Histologic correlates of viral and bacterial infection of the placenta associated with severe morbidity and mortality in the newborn. Hum Pathol. 2004;35:536–545. - PubMed
-
- Gomez R, Romero R, Ghezzi F, Yoon BH, Mazor M, Berry SM. The fetal inflammatory response syndrome. Am J Obstet Gynecol. 1998;179:194–202. - PubMed
-
- Kitchens WH, Chase CM, Uehara S, Cornell LD, Colvin RB, Russell PS, Madsen JC. Macrophage depletion suppresses cardiac allograft vasculopathy in mice. Am J Transplant. 2007;7:2675–2682. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

