Complex reorganization and predominant non-homologous repair following chromosomal breakage in karyotypically balanced germline rearrangements and transgenic integration
- PMID: 22388000
- PMCID: PMC3340016
- DOI: 10.1038/ng.2202
Complex reorganization and predominant non-homologous repair following chromosomal breakage in karyotypically balanced germline rearrangements and transgenic integration
Abstract
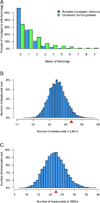
We defined the genetic landscape of balanced chromosomal rearrangements at nucleotide resolution by sequencing 141 breakpoints from cytogenetically interpreted translocations and inversions. We confirm that the recently described phenomenon of 'chromothripsis' (massive chromosomal shattering and reorganization) is not unique to cancer cells but also occurs in the germline, where it can resolve to a relatively balanced state with frequent inversions. We detected a high incidence of complex rearrangements (19.2%) and substantially less reliance on microhomology (31%) than previously observed in benign copy-number variants (CNVs). We compared these results to experimentally generated DNA breakage-repair by sequencing seven transgenic animals, revealing extensive rearrangement of the transgene and host genome with similar complexity to human germline alterations. Inversion was the most common rearrangement, suggesting that a combined mechanism involving template switching and non-homologous repair mediates the formation of balanced complex rearrangements that are viable, stably replicated and transmitted unaltered to subsequent generations.
Figures




References
-
- Iliakis G, et al. Mechanisms of DNA double strand break repair and chromosome aberration formation. Cytogenetic and genome research. 2004;104:14–20. - PubMed
-
- Lee JA, Carvalho CM, Lupski JR. A DNA replication mechanism for generating nonrecurrent rearrangements associated with genomic disorders. Cell. 2007;131:1235–1247. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

