Lysine succinylation and lysine malonylation in histones
- PMID: 22389435
- PMCID: PMC3418837
- DOI: 10.1074/mcp.M111.015875
Lysine succinylation and lysine malonylation in histones
Abstract
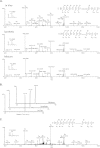
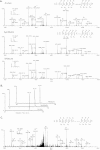
Histone protein post-translational modifications (PTMs) are significant for gene expression and DNA repair. Here we report the identification and validation of a new type of PTM in histones, lysine succinylation. The identified lysine succinylated histone peptides were verified by MS/MS of synthetic peptides, HPLC co-elution, and isotopic labeling. We identified 13, 7, 10, and 7 histone lysine succinylation sites in HeLa, mouse embryonic fibroblast, Drosophila S2, and Saccharomyces cerevisiae cells, respectively. We demonstrated that this histone PTM is present in all eukaryotic cells we examined. Mutagenesis of succinylation sites followed by functional assays implied that histone lysine succinylation can cause unique functional consequences. We also identified one and two histone lysine malonylation sites in HeLa and S. cerevisiae cells, respectively. Our results therefore increase potential combinatorial diversity of histone PTMs and suggest possible new connections between histone biology and metabolism.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

